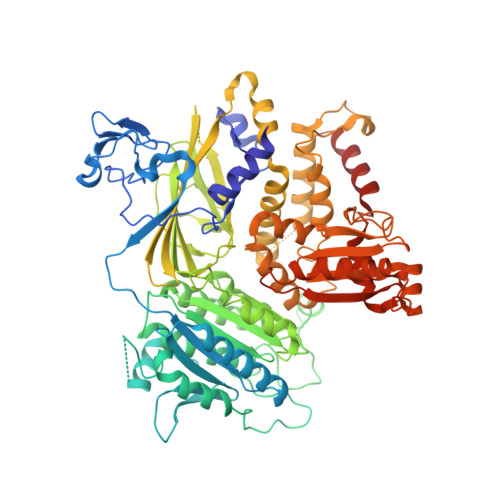
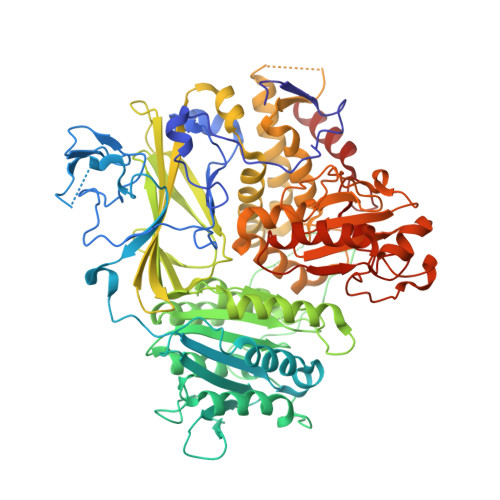
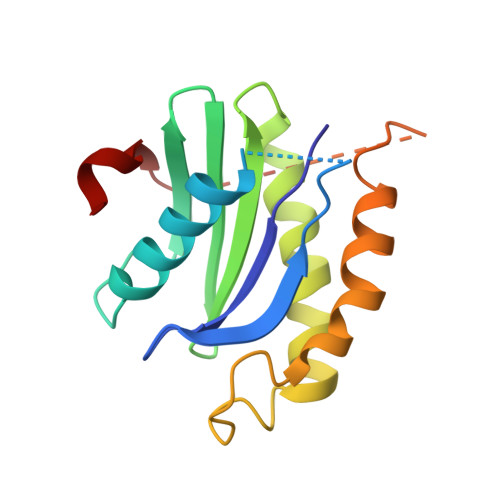
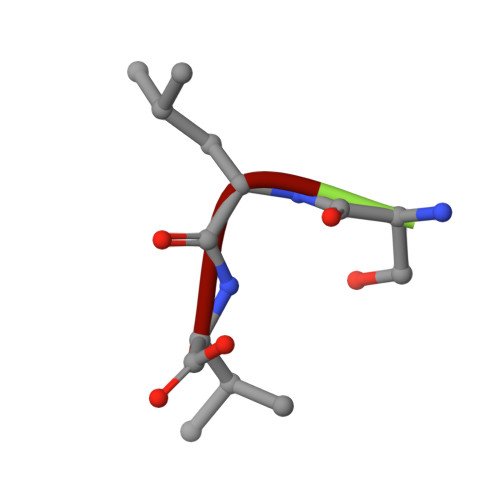
ER retention is imposed by COPII protein sorting and attenuated by 4-phenylbutyrate.
Ma, W., Goldberg, E., Goldberg, J.(2017) Elife 6
- PubMed: 28594326 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.26624
- Primary Citation Related Structures:
5VNE, 5VNF, 5VNG, 5VNH, 5VNI, 5VNJ, 5VNK, 5VNL, 5VNM, 5VNN, 5VNO - PubMed Abstract:
Native cargo proteins exit the endoplasmic reticulum (ER) in COPII-coated vesicles, whereas resident and misfolded proteins are substantially excluded from vesicles by a retention mechanism that remains unresolved. We probed the ER retention process using the proteostasis regulator 4-phenylbutyrate (4-PBA), which we show targets COPII protein to reduce the stringency of retention. 4-PBA competes with p24 proteins to bind COPII. When p24 protein uptake is blocked, COPII vesicles package resident proteins and an ER-trapped mutant LDL receptor. We further show that 4-PBA triggers the secretion of a KDEL-tagged luminal resident, implying that a compromised retention mechanism causes saturation of the KDEL retrieval system. The results indicate that stringent ER retention requires the COPII coat machinery to actively sort biosynthetic cargo from diffusible misfolded and resident ER proteins.
- Structural Biology Program, Memorial Sloan Kettering Cancer Center, New York, United States.
Organizational Affiliation: