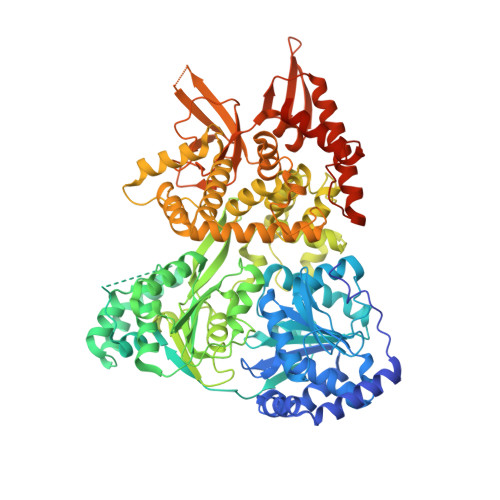
Structural basis of G-quadruplex unfolding by the DEAH/RHA helicase DHX36.
Chen, M.C., Tippana, R., Demeshkina, N.A., Murat, P., Balasubramanian, S., Myong, S., Ferre-D'Amare, A.R.(2018) Nature 558: 465-469
- PubMed: 29899445 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-018-0209-9
- Primary Citation Related Structures:
5VHA, 5VHC, 5VHD, 5VHE - PubMed Abstract:
Guanine-rich nucleic acid sequences challenge the replication, transcription, and translation machinery by spontaneously folding into G-quadruplexes, the unfolding of which requires forces greater than most polymerases can exert 1,2 . Eukaryotic cells contain numerous helicases that can unfold G-quadruplexes 3 . The molecular basis of the recognition and unfolding of G-quadruplexes by helicases remains poorly understood. DHX36 (also known as RHAU and G4R1), a member of the DEAH/RHA family of helicases, binds both DNA and RNA G-quadruplexes with extremely high affinity 4-6 , is consistently found bound to G-quadruplexes in cells 7,8 , and is a major source of G-quadruplex unfolding activity in HeLa cell lysates 6 . DHX36 is a multi-functional helicase that has been implicated in G-quadruplex-mediated transcriptional and post-transcriptional regulation, and is essential for heart development, haematopoiesis, and embryogenesis in mice 9-12 . Here we report the co-crystal structure of bovine DHX36 bound to a DNA with a G-quadruplex and a 3' single-stranded DNA segment. We show that the N-terminal DHX36-specific motif folds into a DNA-binding-induced α-helix that, together with the OB-fold-like subdomain, selectively binds parallel G-quadruplexes. Comparison with unliganded and ATP-analogue-bound DHX36 structures, together with single-molecule fluorescence resonance energy transfer (FRET) analysis, suggests that G-quadruplex binding alone induces rearrangements of the helicase core; by pulling on the single-stranded DNA tail, these rearrangements drive G-quadruplex unfolding one residue at a time.
- Biochemistry and Biophysics Center, National Heart, Lung and Blood Institute, Bethesda, MD, USA.
Organizational Affiliation: