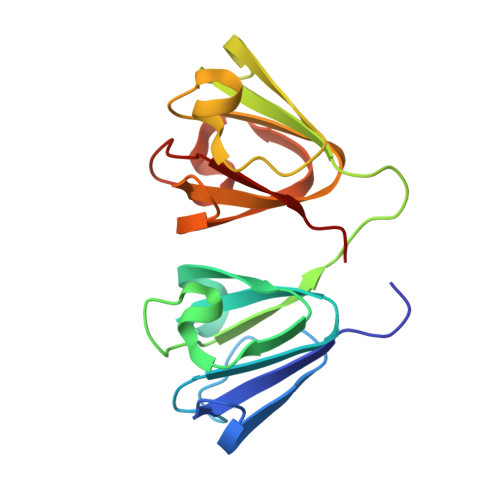
Crystal Structure of Chicken gamma S-Crystallin Reveals Lattice Contacts with Implications for Function in the Lens and the Evolution of the beta gamma-Crystallins.
Sagar, V., Chaturvedi, S.K., Schuck, P., Wistow, G.(2017) Structure 25: 1068-1078.e2
- PubMed: 28648607 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2017.05.015
- Primary Citation Related Structures:
5VH1 - PubMed Abstract:
Previous attempts to crystallize mammalian γS-crystallin were unsuccessful. Native L16 chicken γS crystallized avidly while the Q16 mutant did not. The X-ray structure for chicken γS at 2.3 Å resolution shows the canonical structure of the superfamily plus a well-ordered N arm aligned with a β sheet of a neighboring N domain. L16 is also in a lattice contact, partially shielded from solvent. Unexpectedly, the major lattice contact matches a conserved interface (QR) in the multimeric β-crystallins. QR shows little conservation of residue contacts, except for one between symmetry-related tyrosines, but molecular dipoles for the proteins with QR show striking similarities while other γ-crystallins differ. In γS, QR has few hydrophobic contacts and features a thin layer of tightly bound water. The free energy of QR is slightly repulsive and analytical ultracentrifugation confirms no dimerization in solution. The lattice contacts suggest how γ-crystallins allow close packing without aggregation in the crowded environment of the lens.
- Section on Molecular Structure and Functional Genomics, National Eye Institute, National Institutes of Health, Building 6, Room 106, Bethesda, MD 20892, USA.
Organizational Affiliation: