A Rationally Designed Agonist Defines Subfamily IIIA Abscisic Acid Receptors As Critical Targets for Manipulating Transpiration.
Vaidya, A.S., Peterson, F.C., Yarmolinsky, D., Merilo, E., Verstraeten, I., Park, S.Y., Elzinga, D., Kaundal, A., Helander, J., Lozano-Juste, J., Otani, M., Wu, K., Jensen, D.R., Kollist, H., Volkman, B.F., Cutler, S.R.(2017) ACS Chem Biol 12: 2842-2848
- PubMed: 28949512 Search on PubMed
- DOI: https://doi.org/10.1021/acschembio.7b00650
- Primary Citation Related Structures:
5UR5, 5UR6 - PubMed Abstract:
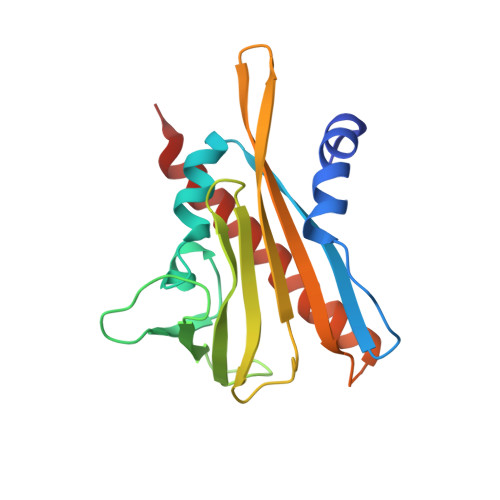
Increasing drought and diminishing freshwater supplies have stimulated interest in developing small molecules that can be used to control transpiration. Receptors for the plant hormone abscisic acid (ABA) have emerged as key targets for this application, because ABA controls the apertures of stomata, which in turn regulate transpiration. Here, we describe the rational design of cyanabactin, an ABA receptor agonist that preferentially activates Pyrabactin Resistance 1 (PYR1) with low nanomolar potency. A 1.63 Å X-ray crystallographic structure of cyanabactin in complex with PYR1 illustrates that cyanabactin's arylnitrile mimics ABA's cyclohexenone oxygen and engages the tryptophan lock, a key component required to stabilize activated receptors. Further, its sulfonamide and 4-methylbenzyl substructures mimic ABA's carboxylate and C6 methyl groups, respectively. Isothermal titration calorimetry measurements show that cyanabactin's compact structure provides ready access to high ligand efficiency on a relatively simple scaffold. Cyanabactin treatments reduce Arabidopsis whole-plant stomatal conductance and activate multiple ABA responses, demonstrating that its in vitro potency translates to ABA-like activity in vivo. Genetic analyses show that the effects of cyanabactin, and the previously identified agonist quinabactin, can be abolished by the genetic removal of PYR1 and PYL1, which form subclade A within the dimeric subfamily III receptors. Thus, cyanabactin is a potent and selective agonist with a wide spectrum of ABA-like activities that defines subfamily IIIA receptors as key target sites for manipulating transpiration.
- Department of Botany and Plant Sciences, Institute of Integrative Genome Biology, University of California , Riverside, California 92521, United States.
Organizational Affiliation: