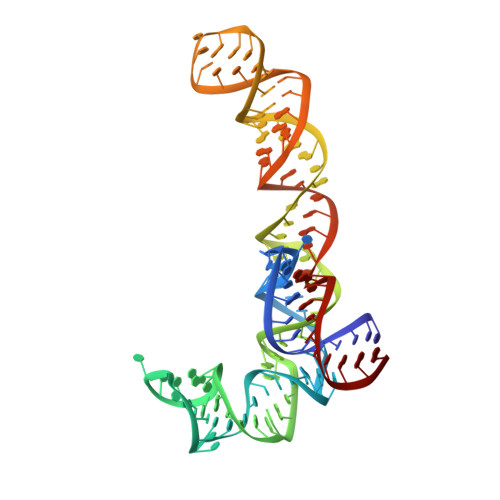
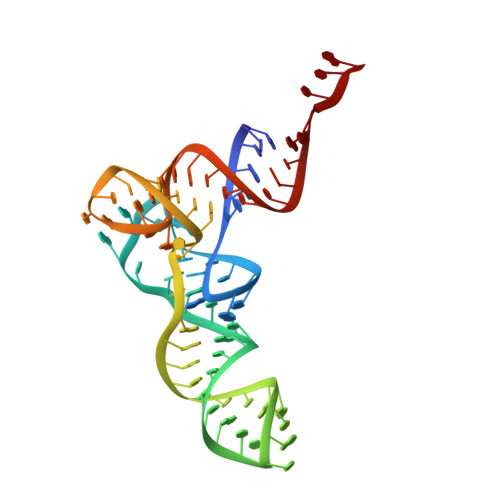
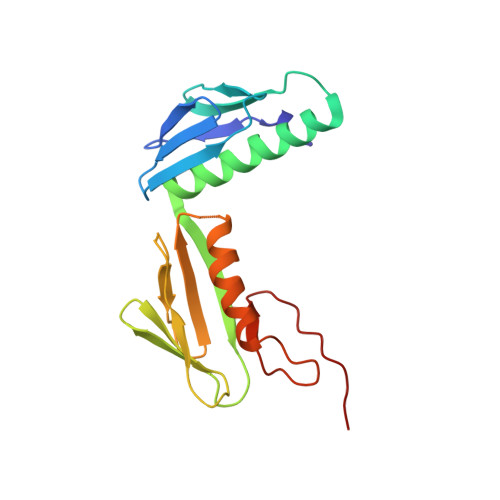
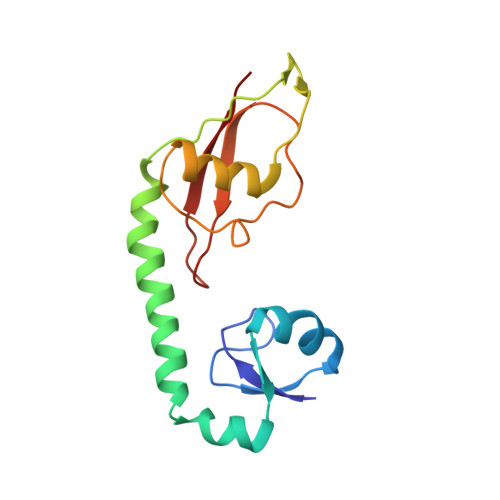
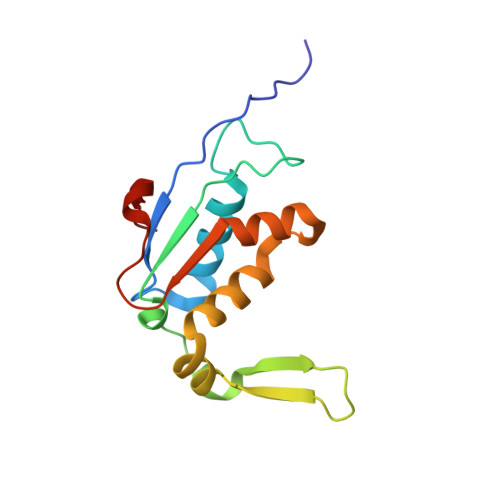
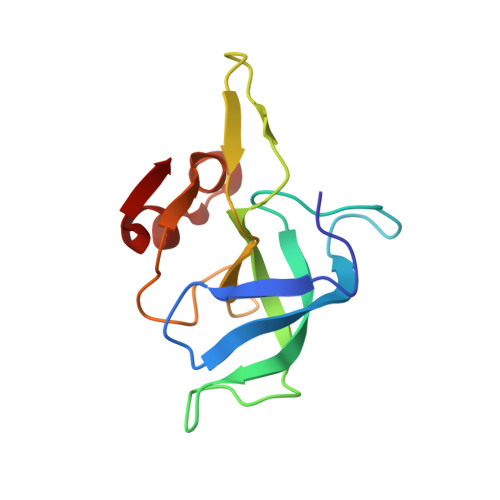
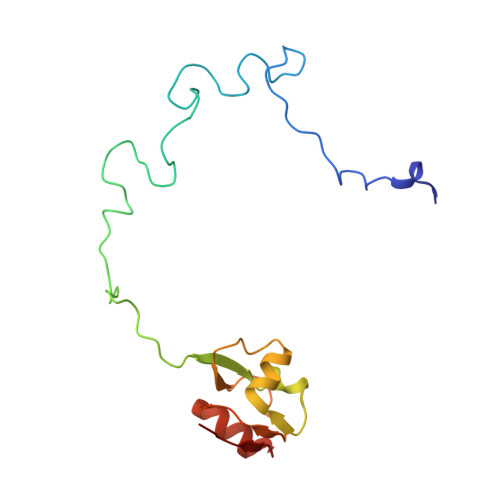
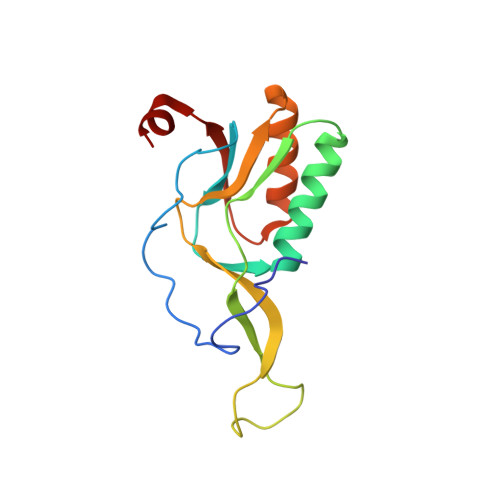
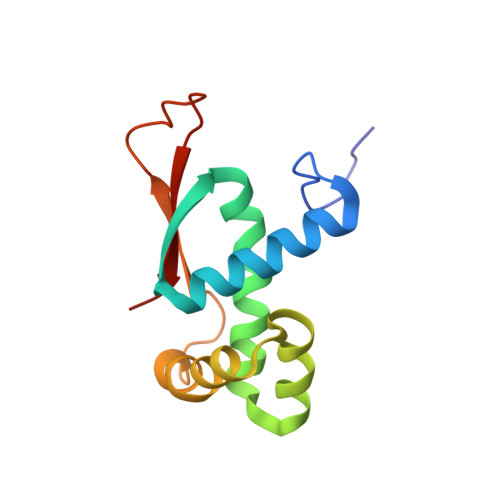
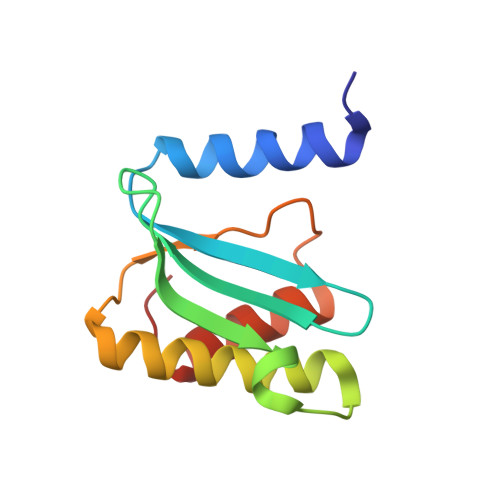
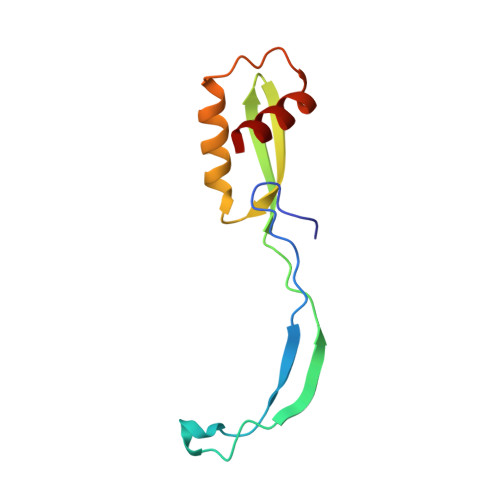
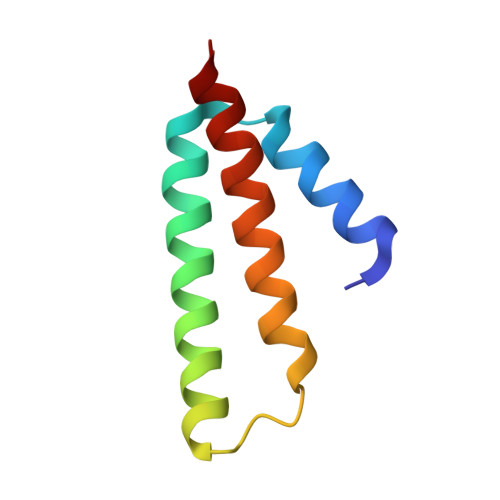
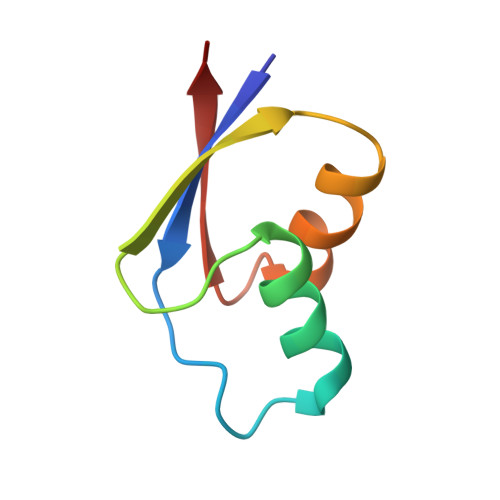
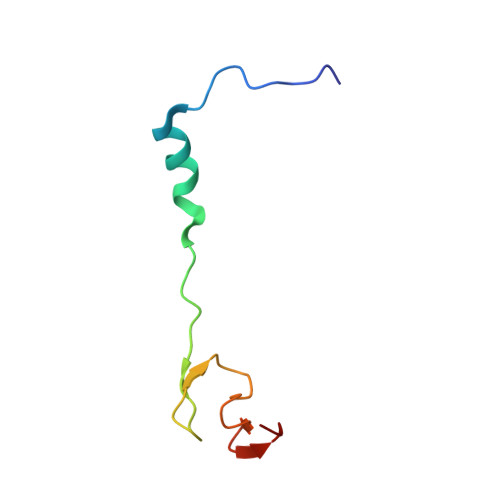
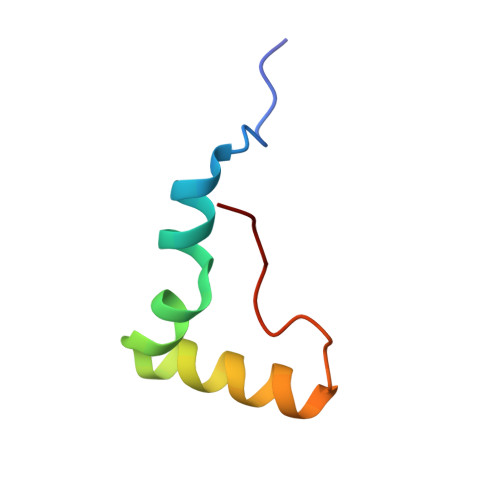
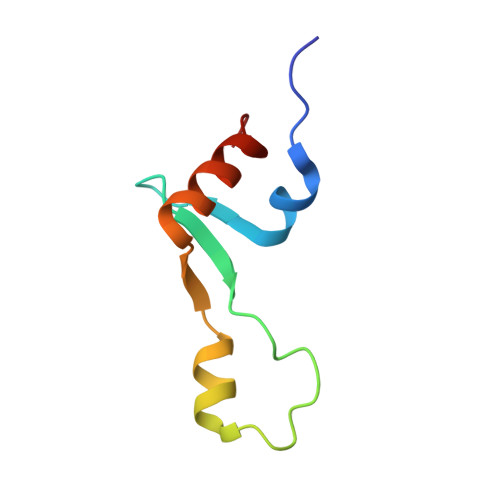
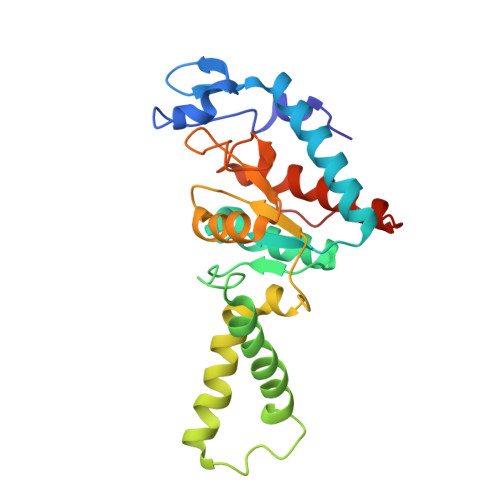
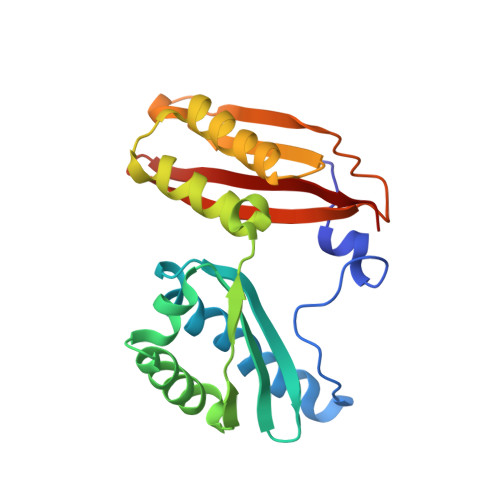
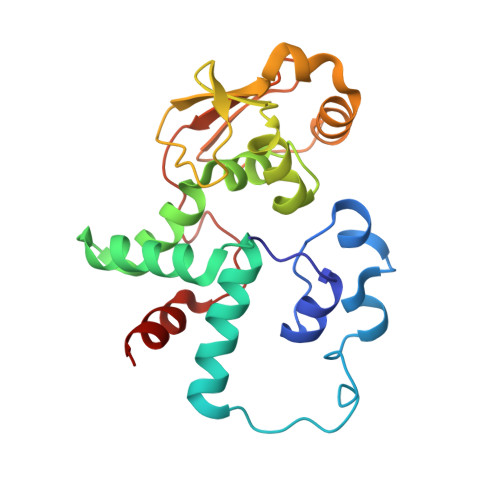
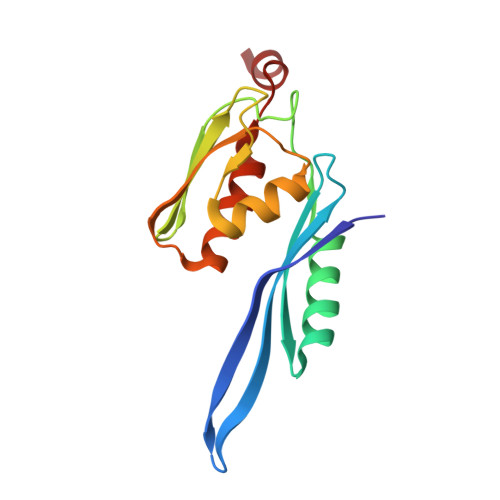
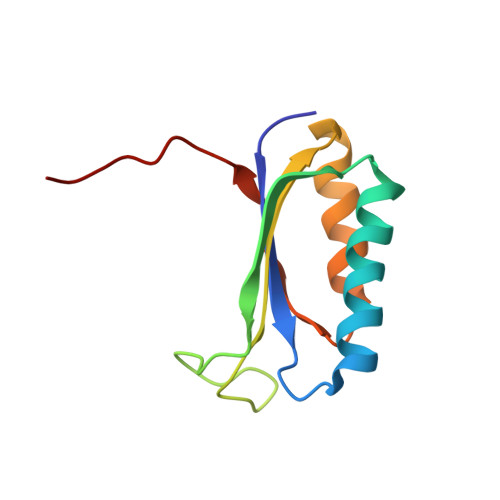
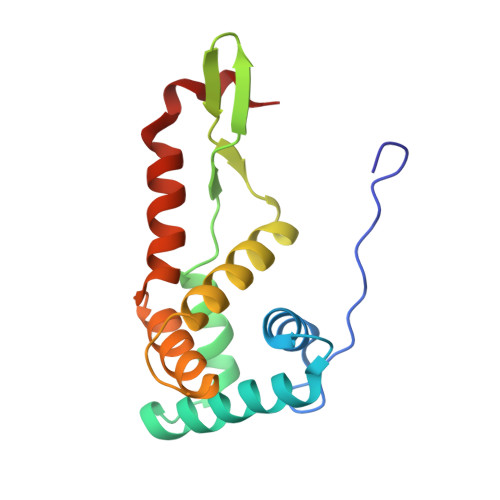
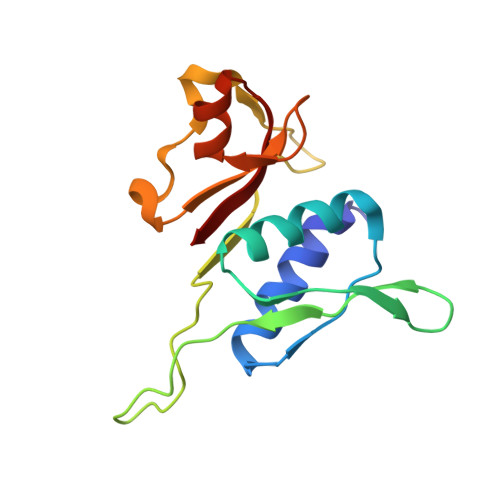
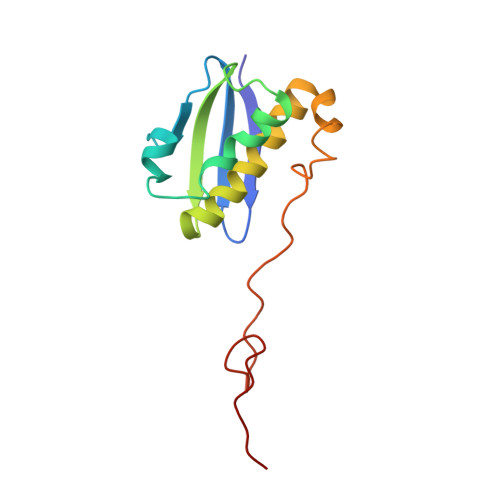
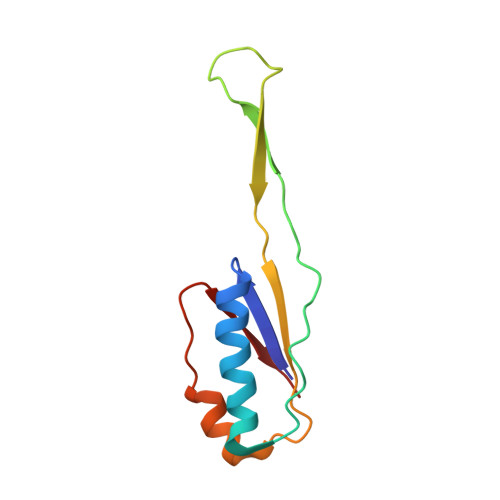
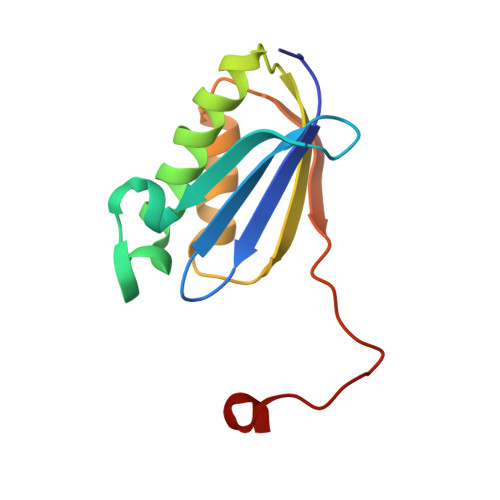
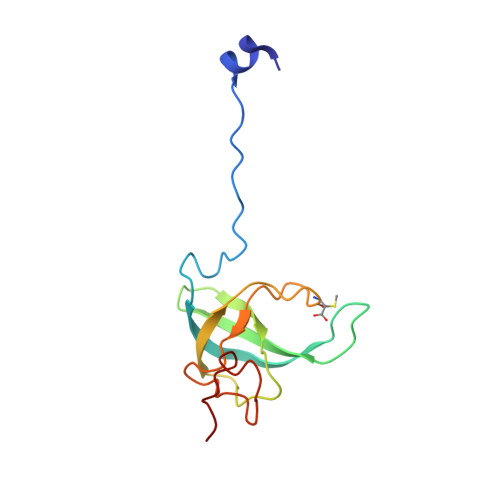
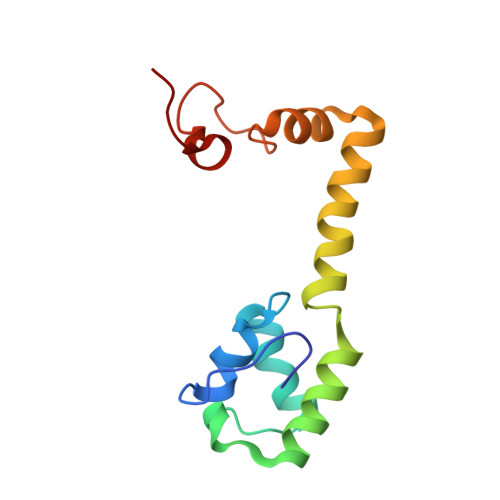
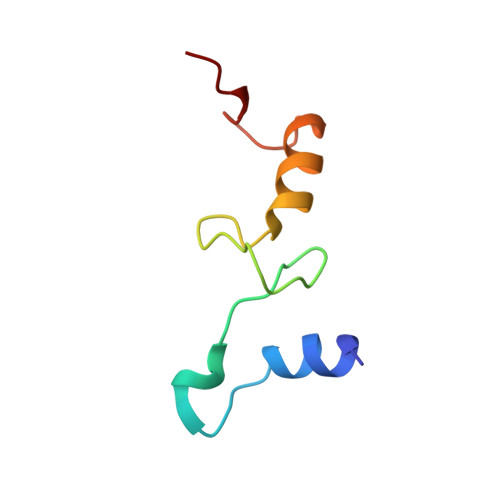
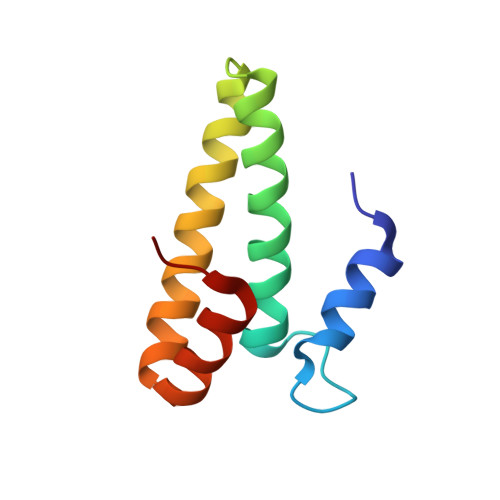
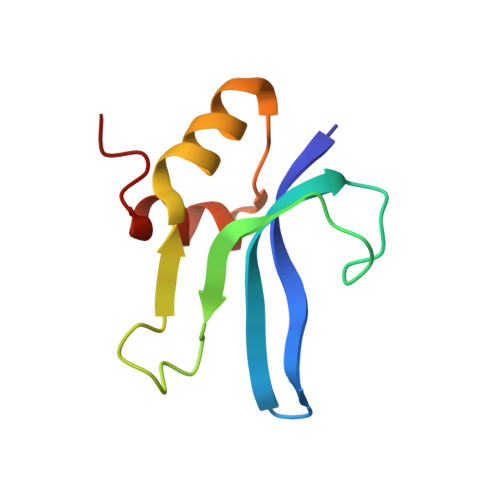
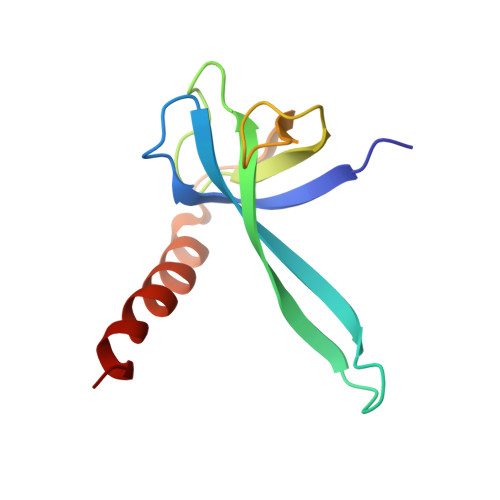
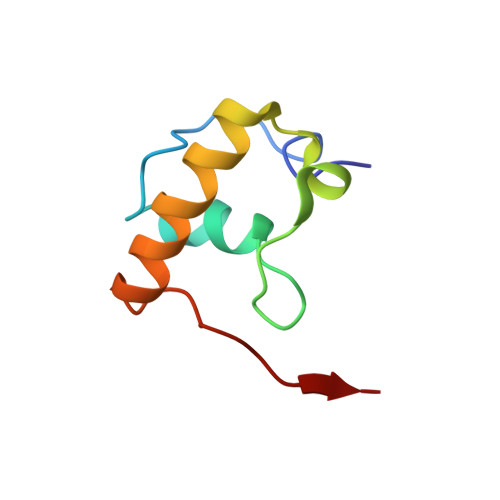
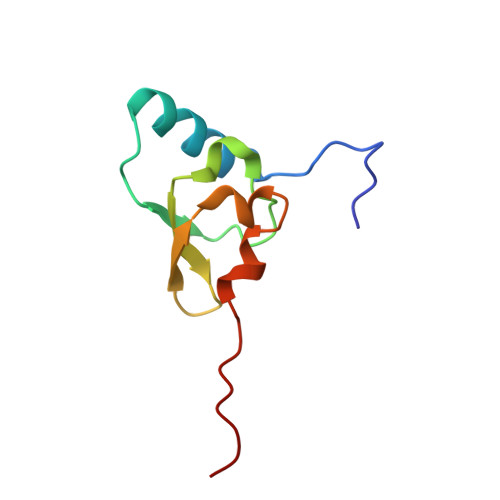
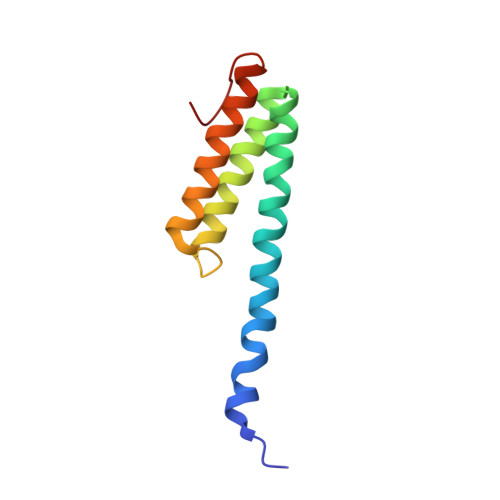
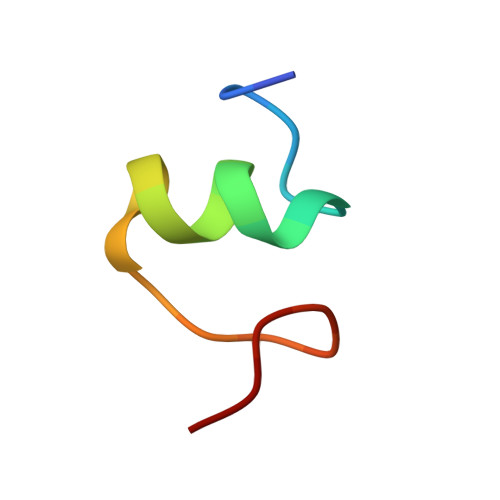
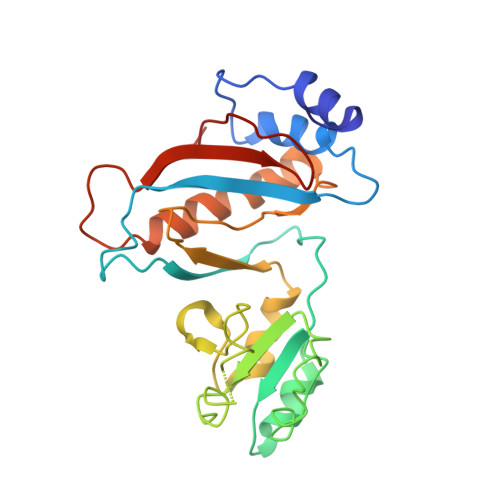
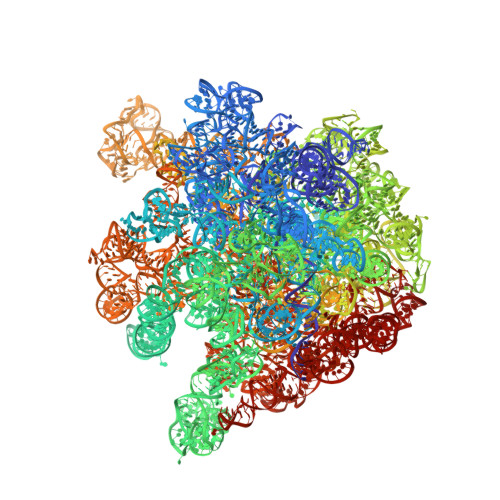
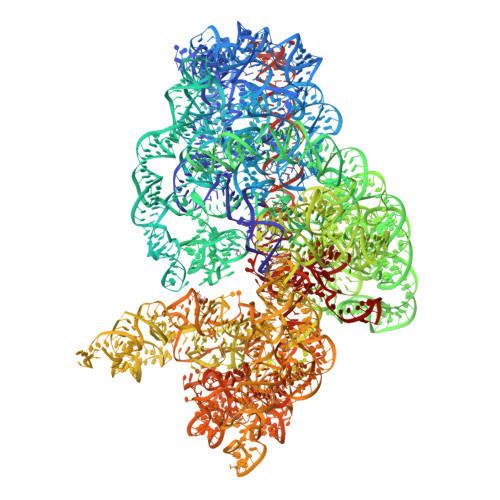
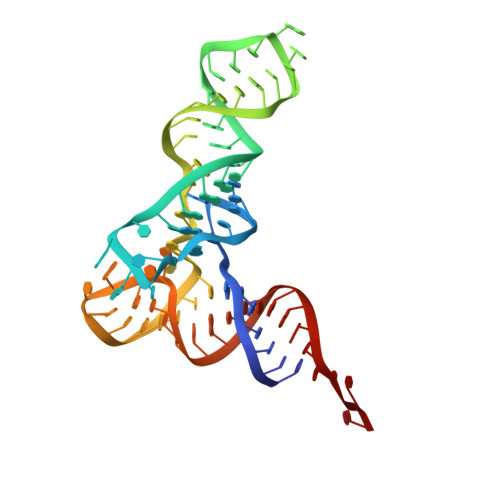
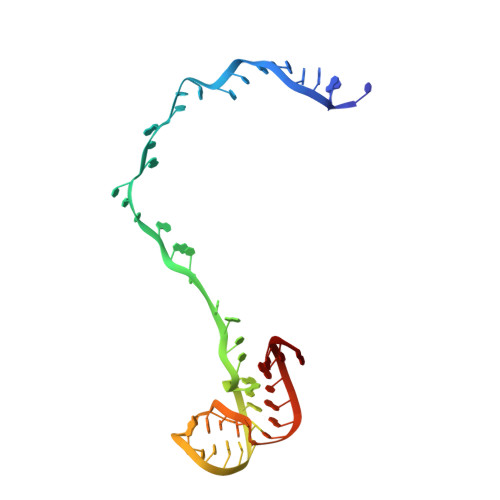
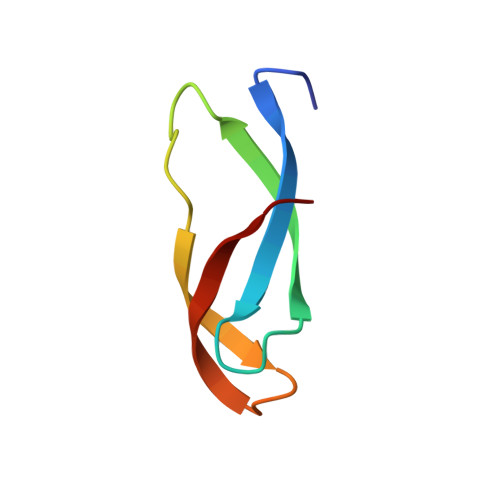Alternative Mode of E-Site tRNA Binding in the Presence of a Downstream mRNA Stem Loop at the Entrance Channel.
Zhang, Y., Hong, S., Ruangprasert, A., Skiniotis, G., Dunham, C.M.(2018) Structure 26: 437-445.e3
- PubMed: 29456023 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2018.01.013
- Primary Citation Related Structures:
5UQ7, 5UQ8 - PubMed Abstract:
Structured mRNAs positioned downstream of the ribosomal decoding center alter gene expression by slowing protein synthesis. Here, we solved the cryo-EM structure of the bacterial ribosome bound to an mRNA containing a 3' stem loop that regulates translation. Unexpectedly, the E-site tRNA adopts two distinct orientations. In the first structure, normal interactions with the 50S and 30S E site are observed. However, in the second structure, although the E-site tRNA makes normal interactions with the 50S E site, its anticodon stem loop moves ∼54 Å away from the 30S E site to interact with the 30S head domain and 50S uL5. This position of the E-site tRNA causes the uL1 stalk to adopt a more open conformation that likely represents an intermediate state during E-site tRNA dissociation. These results suggest that structured mRNAs at the entrance channel restrict 30S subunit movement required during translation to slow E-site tRNA dissociation.
- Life Sciences Institute and Department of Biological Chemistry, University of Michigan, Ann Arbor, MI 48109, USA.
Organizational Affiliation: