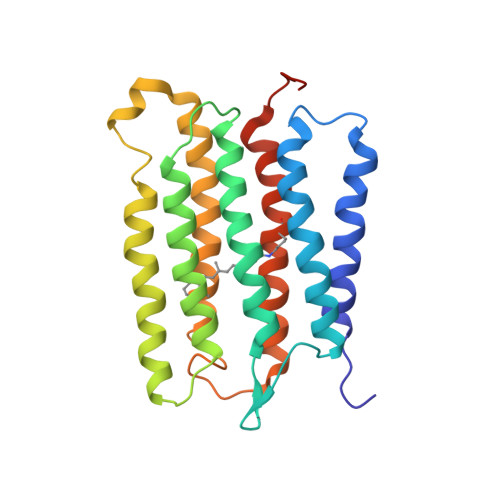
Oligomeric Structure of Anabaena Sensory Rhodopsin in a Lipid Bilayer Environment by Combining Solid-State NMR and Long-range DEER Constraints.
Milikisiyants, S., Wang, S., Munro, R.A., Donohue, M., Ward, M.E., Bolton, D., Brown, L.S., Smirnova, T.I., Ladizhansky, V., Smirnov, A.I.(2017) J Mol Biology 429: 1903-1920
- PubMed: 28501588 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2017.05.005
- Primary Citation Related Structures:
5UK6 - PubMed Abstract:
Oligomerization of membrane proteins is common in nature. Here, we combine spin-labeling double electron-electron resonance (DEER) and solid-state NMR (ssNMR) spectroscopy to refine the structure of an oligomeric integral membrane protein, Anabaena sensory rhodopsin (ASR), reconstituted in a lipid environment. An essential feature of such a combined approach is that it provides structural distance restraints spanning a range of ca 3-60Å while using the same sample preparation (i.e., mutations, paramagnetic labeling, and reconstitution in lipid bilayers) for both ssNMR and DEER. Direct modeling of the multispin effects on DEER signal allowed for the determination of the oligomeric order and for obtaining long-range DEER distance restraints between the ASR trimer subunits that were used to refine the ssNMR structure of ASR. The improved structure of the ASR trimer revealed a more compact packing of helices and side chains at the intermonomer interface, compared to the structure determined using the ssNMR data alone. The extent of the refinement is significant when compared with typical helix movements observed for the active states of homologous proteins. Our combined approach of using complementary DEER and NMR measurements for the determination of oligomeric structures would be widely applicable to membrane proteins where paramagnetic tags can be introduced. Such a method could be used to study the effects of the lipid membrane composition on protein oligomerization and to observe structural changes in protein oligomers upon drug, substrate, and co-factor binding.
- Department of Chemistry, College of Sciences, North Carolina State University, 2620 Yarbrough Dive, Raleigh, NC 27695-8204, USA.
Organizational Affiliation: