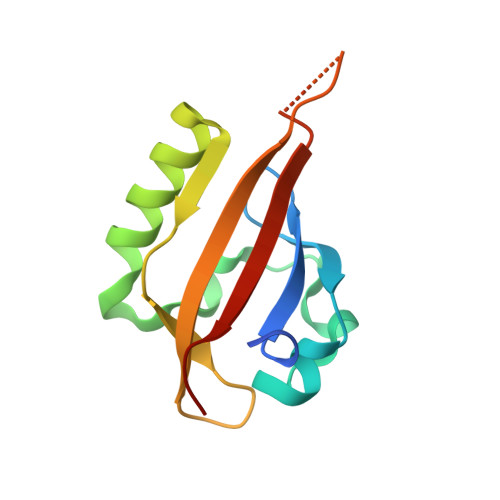
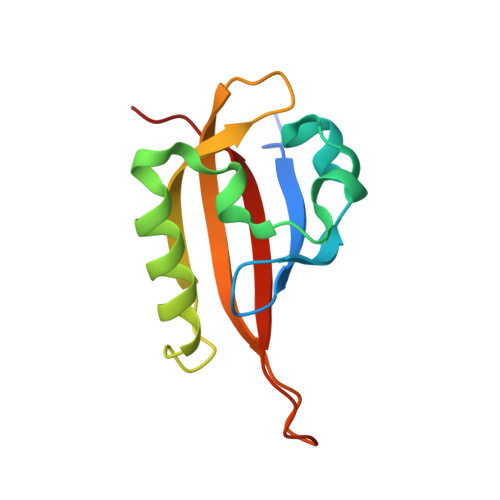
On-target efficacy of a HIF-2 alpha antagonist in preclinical kidney cancer models.
Cho, H., Du, X., Rizzi, J.P., Liberzon, E., Chakraborty, A.A., Gao, W., Carvo, I., Signoretti, S., Bruick, R.K., Josey, J.A., Wallace, E.M., Kaelin, W.G.(2016) Nature 539: 107-111
- PubMed: 27595393 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature19795
- Primary Citation Related Structures:
5UFP - PubMed Abstract:
Clear cell renal cell carcinoma, the most common form of kidney cancer, is usually linked to inactivation of the pVHL tumour suppressor protein and consequent accumulation of the HIF-2α transcription factor (also known as EPAS1). Here we show that a small molecule (PT2399) that directly inhibits HIF-2α causes tumour regression in preclinical mouse models of primary and metastatic pVHL-defective clear cell renal cell carcinoma in an on-target fashion. pVHL-defective clear cell renal cell carcinoma cell lines display unexpectedly variable sensitivity to PT2399, however, suggesting the need for predictive biomarkers to be developed to use this approach optimally in the clinic.
- Department of Medical Oncology, Dana-Farber Cancer Institute and Brigham and Women's Hospital, Harvard Medical School, Boston, Massachusetts 02215, USA.
Organizational Affiliation: