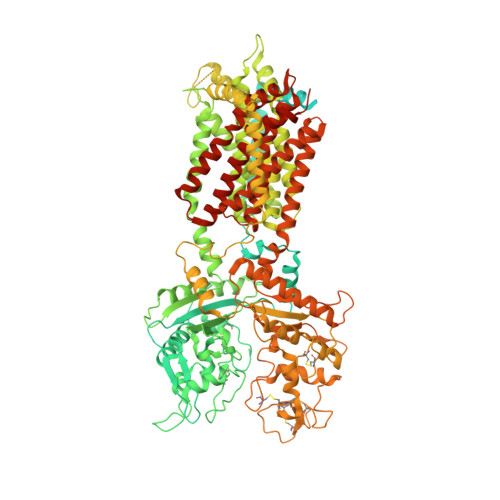
3.3 angstrom structure of Niemann-Pick C1 protein reveals insights into the function of the C-terminal luminal domain in cholesterol transport.
Li, X., Lu, F., Trinh, M.N., Schmiege, P., Seemann, J., Wang, J., Blobel, G.(2017) Proc Natl Acad Sci U S A 114: 9116-9121
- PubMed: 28784760 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1711716114
- Primary Citation Related Structures:
5U73, 5U74 - PubMed Abstract:
Niemann-Pick C1 (NPC1) and NPC2 proteins are indispensable for the export of LDL-derived cholesterol from late endosomes. Mutations in these proteins result in Niemann-Pick type C disease, a lysosomal storage disease. Despite recent reports of the NPC1 structure depicting its overall architecture, the function of its C-terminal luminal domain (CTD) remains poorly understood even though 45% of NPC disease-causing mutations are in this domain. Here, we report a crystal structure at 3.3 Å resolution of NPC1* (residues 314-1,278), which-in contrast to previous lower resolution structures-features the entire CTD well resolved. Notably, all eight cysteines of the CTD form four disulfide bonds, one of which (C909-C914) enforces a specific loop that in turn mediates an interaction with a loop of the N-terminal domain (NTD). Importantly, this loop and its interaction with the NTD were not observed in any previous structures due to the lower resolution. Our mutagenesis experiments highlight the physiological relevance of the CTD-NTD interaction, which might function to keep the NTD in the proper orientation for receiving cholesterol from NPC2. Additionally, this structure allows us to more precisely map all of the disease-causing mutations, allowing future molecular insights into the pathogenesis of NPC disease.
- Laboratory of Cell Biology, The Rockefeller University, New York, NY 10065; xli05@rockefeller.edu blobel@rockefeller.edu.
Organizational Affiliation: