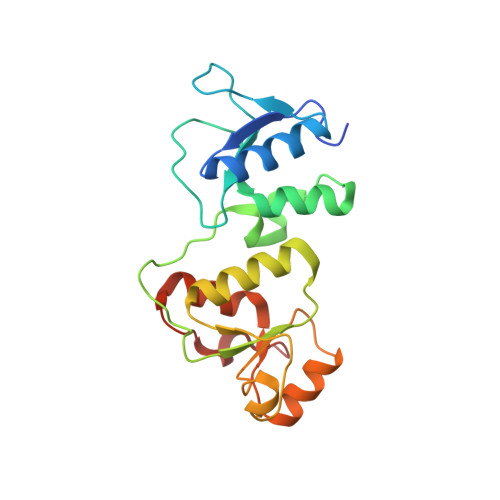
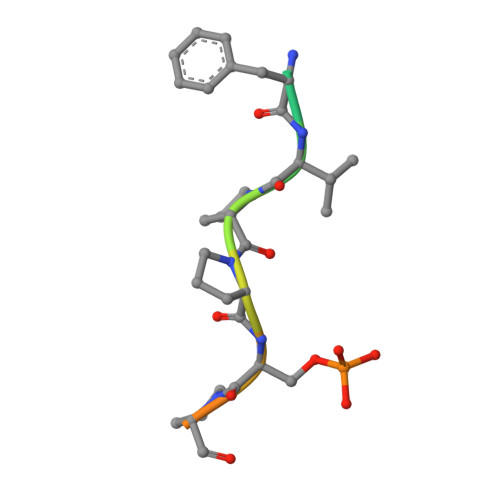
Structural Insight into BLM Recognition by TopBP1.
Sun, L., Huang, Y., Edwards, R.A., Yang, S., Blackford, A.N., Niedzwiedz, W., Glover, J.N.M.(2017) Structure 25: 1582-1588.e3
- PubMed: 28919440 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2017.08.005
- Primary Citation Related Structures:
5U6K - PubMed Abstract:
Topoisomerase IIβ binding protein 1 (TopBP1) is a critical protein-protein interaction hub in DNA replication checkpoint control. It was proposed that TopBP1 BRCT5 interacts with Bloom syndrome helicase (BLM) to regulate genome stability through either phospho-Ser304 or phospho-Ser338 of BLM. Here we show that TopBP1 BRCT5 specifically interacts with the BLM region surrounding pSer304, not pSer338. Our crystal structure of TopBP1 BRCT4/5 bound to BLM reveals recognition of pSer304 by a conserved pSer-binding pocket, and interactions between an FVPP motif N-terminal to pSer304 and a hydrophobic groove on BRCT5. This interaction utilizes the same surface of BRCT5 that recognizes the DNA damage mediator, MDC1; however the binding orientations of MDC1 and BLM are reversed. While the MDC1 interactions are largely electrostatic, the interaction with BLM has higher affinity and relies on a mix of electrostatics and hydrophobicity. We suggest that similar evolutionarily conserved interactions may govern interactions between TopBP1 and 53BP1.
- Department of Biochemistry, University of Alberta, Edmonton, AB T6G 2H7, Canada.
Organizational Affiliation: