Crystal Structure Of I83E Meditope-Enabled Trastuzumab With Azido-PEG3-Meditope
Williams, J.C., Bzymek, K.P., Pucket, J., Avery, K.A., Ma, Y., Xie, J., Zer, C., Horne, D.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Light Chain | 214 | Homo sapiens | Mutation(s): 0 | 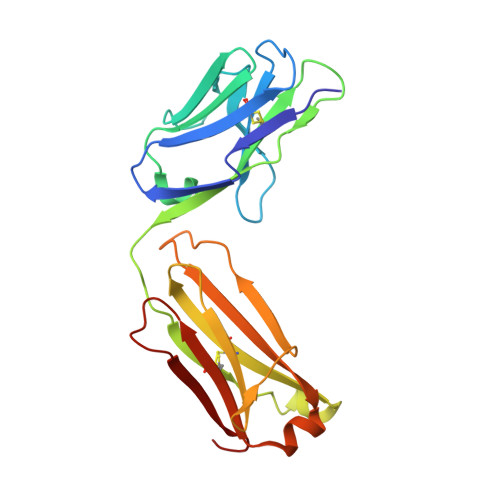 | |
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Heavy Chain | 223 | Homo sapiens | Mutation(s): 0 | 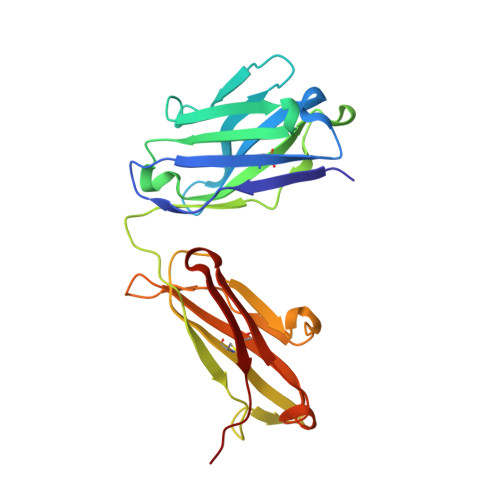 | |
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Protein L | C [auth E] | 63 | Finegoldia magna | Mutation(s): 5 | 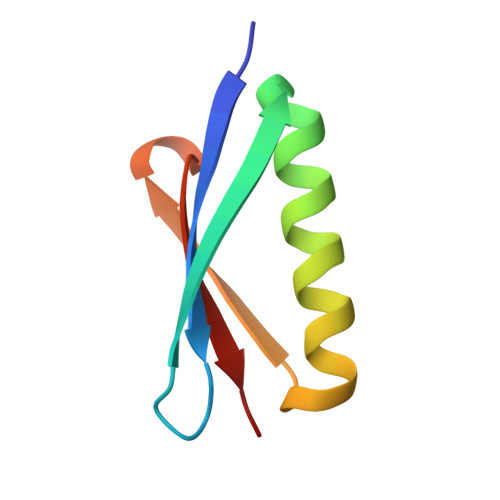 |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q51918 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 4 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Immunoglobulin G binding protein A | D [auth C] | 54 | Staphylococcus aureus | Mutation(s): 0 Gene Names: spa | 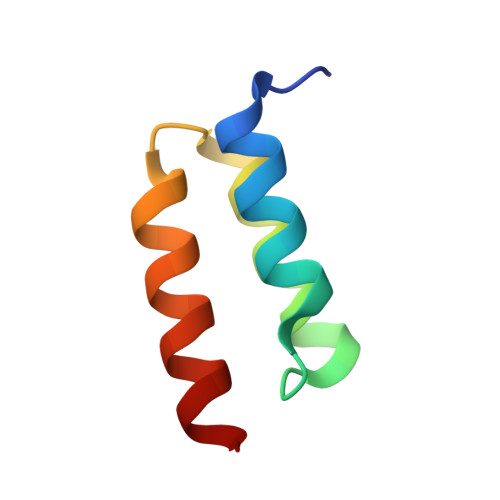 |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P02976 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 5 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| meditope peptide | E [auth D] | 14 | synthetic construct | Mutation(s): 0 | 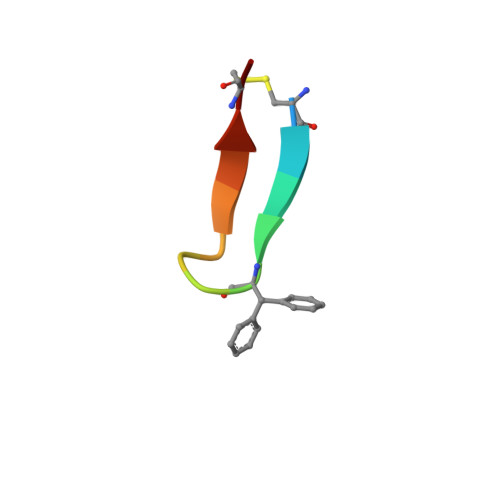 |
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| MRY Download:Ideal Coordinates CCD File | F [auth B] | MESO-ERYTHRITOL C4 H10 O4 UNXHWFMMPAWVPI-ZXZARUISSA-N |  | ||
| Modified Residues 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| 2GX Query on 2GX | E [auth D] | L-PEPTIDE LINKING | C15 H15 N O2 |  | PHE |
| 562 Query on 562 | E [auth D] | L-PEPTIDE LINKING | C12 H26 N7 O4 |  | -- |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 53.25 | α = 90 |
| b = 105.17 | β = 90 |
| c = 117.07 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| XDS | data reduction |
| XSCALE | data scaling |
| MOLREP | phasing |