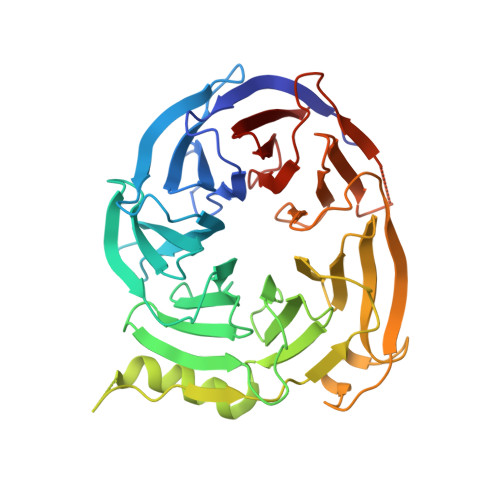
Structure-Guided Design of EED Binders Allosterically Inhibiting the Epigenetic Polycomb Repressive Complex 2 (PRC2) Methyltransferase.
Lingel, A., Sendzik, M., Huang, Y., Shultz, M.D., Cantwell, J., Dillon, M.P., Fu, X., Fuller, J., Gabriel, T., Gu, J., Jiang, X., Li, L., Liang, F., McKenna, M., Qi, W., Rao, W., Sheng, X., Shu, W., Sutton, J., Taft, B., Wang, L., Zeng, J., Zhang, H., Zhang, M., Zhao, K., Lindvall, M., Bussiere, D.E.(2017) J Med Chem 60: 415-427
- PubMed: 27992714 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.6b01473
- Primary Citation Related Structures:
5U5H, 5U5K, 5U5T, 5U62 - PubMed Abstract:
PRC2 is a multisubunit methyltransferase involved in epigenetic regulation of early embryonic development and cell growth. The catalytic subunit EZH2 methylates primarily lysine 27 of histone H3, leading to chromatin compaction and repression of tumor suppressor genes. Inhibiting this activity by small molecules targeting EZH2 was shown to result in antitumor efficacy. Here, we describe the optimization of a chemical series representing a new class of PRC2 inhibitors which acts allosterically via the trimethyllysine pocket of the noncatalytic EED subunit. Deconstruction of a larger and complex screening hit to a simple fragment-sized molecule followed by structure-guided regrowth and careful property modulation were employed to yield compounds which achieve submicromolar inhibition in functional assays and cellular activity. The resulting molecules can serve as a simplified entry point for lead optimization and can be utilized to study this new mechanism of PRC2 inhibition and the associated biology in detail.
- Novartis Institutes for BioMedical Research , 5300 Chiron Way, Emeryville, California 94608, United States.
Organizational Affiliation: