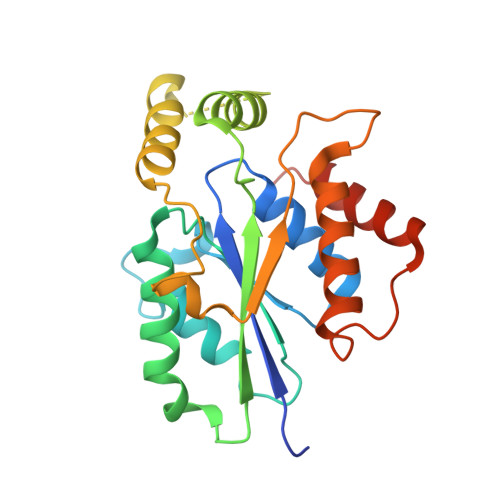
Structural basis for the GTP specificity of the RNA kinase domain of fungal tRNA ligase.
Remus, B.S., Goldgur, Y., Shuman, S.(2017) Nucleic Acids Res 45: 12945-12953
- PubMed: 29165709 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkx1159
- Primary Citation Related Structures:
5U32 - PubMed Abstract:
Fungal tRNA ligase (Trl1) is an essential enzyme that repairs RNA breaks with 2',3'-cyclic-PO4 and 5'-OH ends inflicted during tRNA splicing and non-canonical mRNA splicing in the fungal unfolded protein response. Trl1 is composed of C-terminal cyclic phosphodiesterase and central polynucleotide kinase domains that heal the broken ends to generate the 3'-OH,2'-PO4 and 5'-PO4 termini required for sealing by an N-terminal ligase domain. Trl1 enzymes are found in all human fungal pathogens and are promising targets for antifungal drug discovery because their domain compositions and biochemical mechanisms are unique compared to the mammalian RtcB-type tRNA splicing enzyme. A distinctive feature of Trl1 is its preferential use of GTP as phosphate donor for the RNA kinase reaction. Here we report the 2.2 Å crystal structure of the kinase domain of Trl1 from the fungal pathogen Candida albicans with GDP and Mg2+ in the active site. The P-loop phosphotransferase fold of the kinase is embellished by a unique 'G-loop' element that accounts for guanine nucleotide specificity. Mutations of amino acids that contact the guanine nucleobase efface kinase activity in vitro and Trl1 function in vivo. Our findings fortify the case for the Trl1 kinase as an antifungal target.
- Molecular Biology Program, Sloan-Kettering Institute, New York, NY 10065, USA.
Organizational Affiliation: