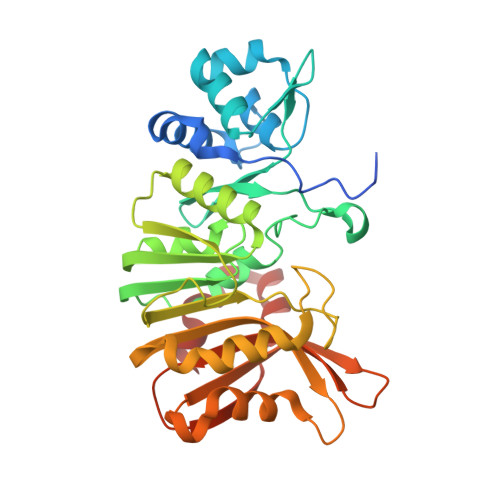
Structural Basis of the Selectivity of GenN, an Aminoglycoside N-Methyltransferase Involved in Gentamicin Biosynthesis.
Bury, P.D.S., Huang, F., Li, S., Sun, Y., Leadlay, P.F., Dias, M.V.B.(2017) ACS Chem Biol 12: 2779-2787
- PubMed: 28876898 Search on PubMed
- DOI: https://doi.org/10.1021/acschembio.7b00466
- Primary Citation Related Structures:
5TYQ, 5U0N, 5U0T, 5U18, 5U19, 5U1E, 5U1I, 5U4T - PubMed Abstract:
Gentamicins are heavily methylated, clinically valuable pseudotrisaccharide antibiotics produced by Micromonospora echinospora. GenN has been characterized as an S-adenosyl-l-methionine-dependent methyltransferase with low sequence similarity to other enzymes. It is responsible for the 3″-N-methylation of 3″-dehydro-3″-amino-gentamicin A2, an essential modification of ring III in the biosynthetic pathway to the gentamicin C complex. Purified recombinant GenN also efficiently catalyzes 3″-N-methylation of related aminoglycosides kanamycin B and tobramycin, which both contain an additional hydroxymethyl group at the C5″ position in ring III. We have obtained eight cocrystal structures of GenN, at a resolution of 2.2 Å or better, including the binary complex of GenN and S-adenosyl-l-homocysteine (SAH) and the ternary complexes of GenN, SAH, and several aminoglycosides. The GenN structure reveals several features not observed in any other N-methyltransferase that fit it for its role in gentamicin biosynthesis. These include a novel N-terminal domain that might be involved in protein:protein interaction with upstream enzymes of the gentamicin X2 biosynthesis and two long loops that are involved in aminoglycoside substrate recognition. In addition, the analysis of structures of GenN in complex with different ligands, supported by the results of active site mutagenesis, has allowed us to propose a catalytic mechanism and has revealed the structural basis for the surprising ability of native GenN to act on these alternative substrates.
- Department of Microbiology, Institute of Biomedical Science, University of São Paulo , São Paulo, Brazil.
Organizational Affiliation: