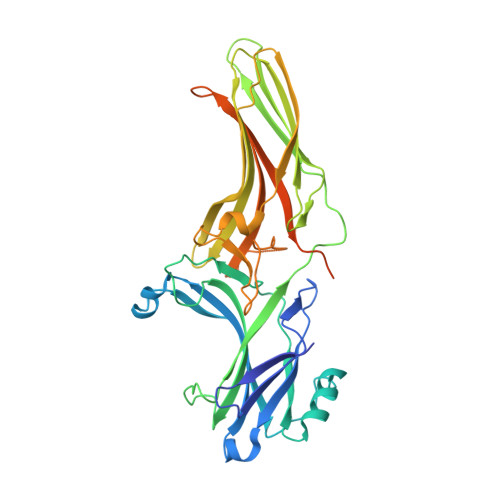
Structural basis of arrestin-3 activation and signaling.
Chen, Q., Perry, N.A., Vishnivetskiy, S.A., Berndt, S., Gilbert, N.C., Zhuo, Y., Singh, P.K., Tholen, J., Ohi, M.D., Gurevich, E.V., Brautigam, C.A., Klug, C.S., Gurevich, V.V., Iverson, T.M.(2017) Nat Commun 8: 1427-1427
- PubMed: 29127291 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-017-01218-8
- Primary Citation Related Structures:
5TV1 - PubMed Abstract:
A unique aspect of arrestin-3 is its ability to support both receptor-dependent and receptor-independent signaling. Here, we show that inositol hexakisphosphate (IP 6 ) is a non-receptor activator of arrestin-3 and report the structure of IP 6 -activated arrestin-3 at 2.4-Å resolution. IP 6 -activated arrestin-3 exhibits an inter-domain twist and a displaced C-tail, hallmarks of active arrestin. IP 6 binds to the arrestin phosphate sensor, and is stabilized by trimerization. Analysis of the trimerization surface, which is also the receptor-binding surface, suggests a feature called the finger loop as a key region of the activation sensor. We show that finger loop helicity and flexibility may underlie coupling to hundreds of diverse receptors and also promote arrestin-3 activation by IP 6 . Importantly, we show that effector-binding sites on arrestins have distinct conformations in the basal and activated states, acting as switch regions. These switch regions may work with the inter-domain twist to initiate and direct arrestin-mediated signaling.
- Department of Pharmacology, Vanderbilt University, Nashville, TN, 37232, USA.
Organizational Affiliation: