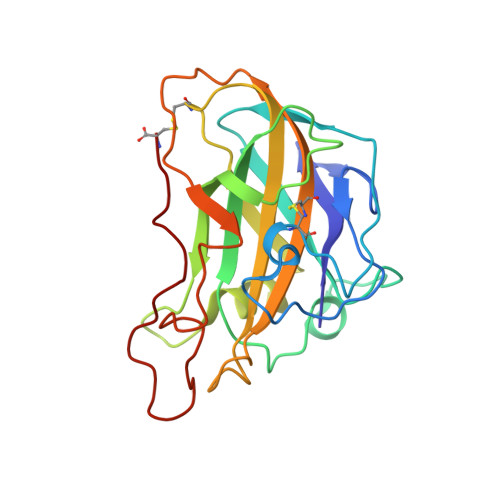
Crystallization of a fungal lytic polysaccharide monooxygenase expressed from glycoengineered Pichia pastoris for X-ray and neutron diffraction.
O'Dell, W.B., Swartz, P.D., Weiss, K.L., Meilleur, F.(2017) Acta Crystallogr F Struct Biol Commun 73: 70-78
- PubMed: 28177316 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X16020318
- Primary Citation Related Structures:
5TKF - PubMed Abstract:
Lytic polysaccharide monooxygenases (LPMOs) are carbohydrate-disrupting enzymes secreted by bacteria and fungi that break glycosidic bonds via an oxidative mechanism. Fungal LPMOs typically act on cellulose and can enhance the efficiency of cellulose-hydrolyzing enzymes that release soluble sugars for bioethanol production or other industrial uses. The enzyme PMO-2 from Neurospora crassa (NcPMO-2) was heterologously expressed in Pichia pastoris to facilitate crystallographic studies of the fungal LPMO mechanism. Diffraction resolution and crystal morphology were improved by expressing NcPMO-2 from a glycoengineered strain of P. pastoris and by the use of crystal seeding methods, respectively. These improvements resulted in high-resolution (1.20 Å) X-ray diffraction data collection at 100 K and the production of a large NcPMO-2 crystal suitable for room-temperature neutron diffraction data collection to 2.12 Å resolution.
- Department of Molecular and Structural Biochemistry, North Carolina State University, Campus Box 7622, Raleigh, NC 27695, USA.
Organizational Affiliation: