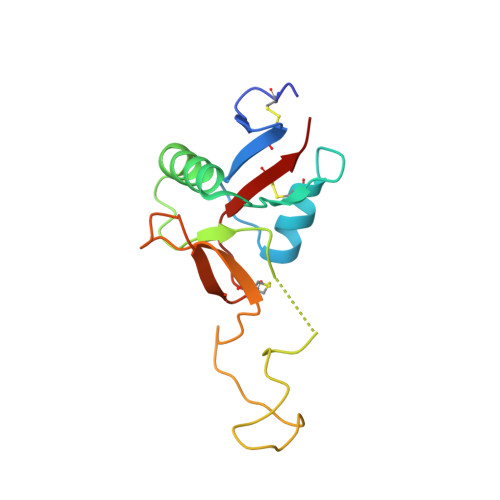
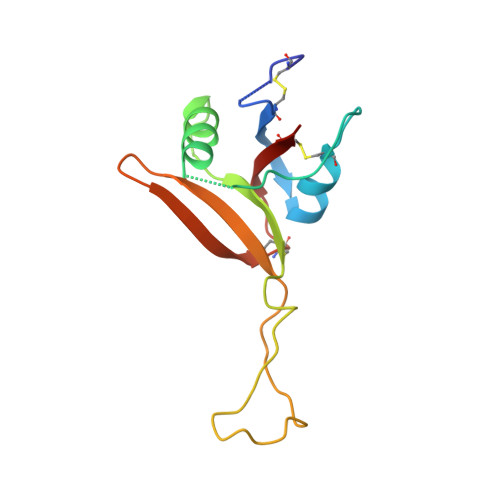
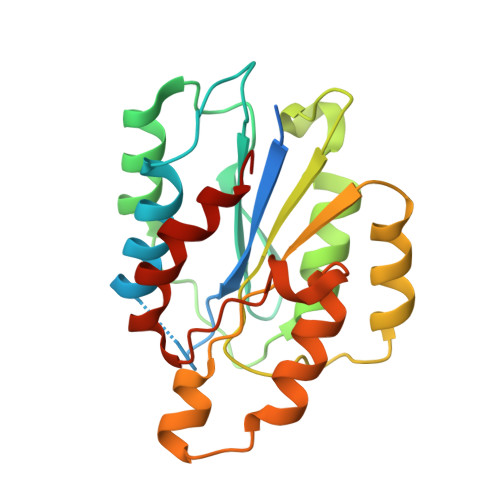
Dramatic and concerted conformational changes enable rhodocetin to block alpha 2 beta 1 integrin selectively.
Eble, J.A., McDougall, M., Orriss, G.L., Niland, S., Johanningmeier, B., Pohlentz, G., Meier, M., Karrasch, S., Estevao-Costa, M.I., Martins Lima, A., Stetefeld, J.(2017) PLoS Biol 15: e2001492-e2001492
- PubMed: 28704364 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pbio.2001492
- Primary Citation Related Structures:
5THP - PubMed Abstract:
The collagen binding integrin α2β1 plays a crucial role in hemostasis, fibrosis, and cancer progression amongst others. It is specifically inhibited by rhodocetin (RC), a C-type lectin-related protein (CLRP) found in Malayan pit viper (Calloselasma rhodostoma) venom. The structure of RC alone reveals a heterotetramer arranged as an αβ and γδ subunit in a cruciform shape. RC specifically binds to the collagen binding A-domain of the integrin α2 subunit, thereby blocking collagen-induced platelet aggregation. However, until now, the molecular basis for this interaction has remained unclear. Here, we present the molecular structure of the RCγδ-α2A complex solved to 3.0 Å resolution. Our findings show that RC undergoes a dramatic structural reorganization upon binding to α2β1 integrin. Besides the release of the nonbinding RCαβ tandem, the RCγ subunit interacts with loop 2 of the α2A domain as result of a dramatic conformational change. The RCδ subunit contacts the integrin α2A domain in the "closed" conformation through its helix C. Combined with epitope-mapped antibodies, conformationally locked α2A domain mutants, point mutations within the α2A loop 2, and chemical modifications of the purified toxin protein, this molecular structure of RCγδ-α2A complex explains the inhibitory mechanism and specificity of RC for α2β1 integrin.
- Institute of Physiological Chemistry and Pathobiochemistry, University of Münster, Münster, Germany.
Organizational Affiliation: