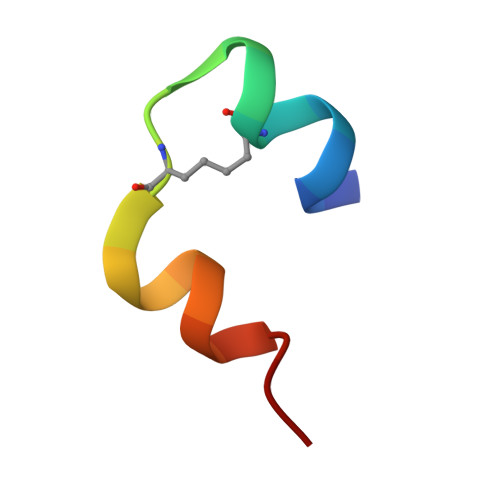
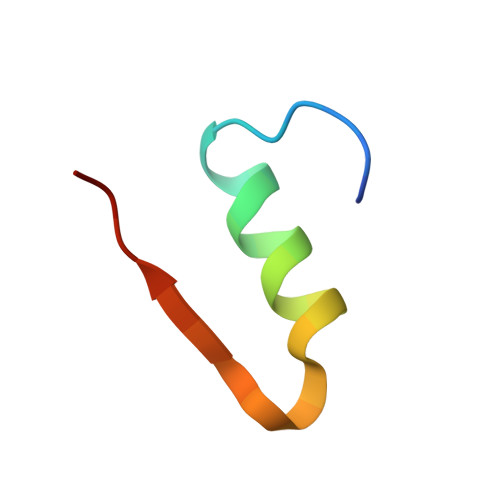
Insulin in motion: The A6-A11 disulfide bond allosterically modulates structural transitions required for insulin activity.
van Lierop, B., Ong, S.C., Belgi, A., Delaine, C., Andrikopoulos, S., Haworth, N.L., Menting, J.G., Lawrence, M.C., Robinson, A.J., Forbes, B.E.(2017) Sci Rep 7: 17239-17239
- PubMed: 29222417 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-017-16876-3
- Primary Citation Related Structures:
5T7R - PubMed Abstract:
The structural transitions required for insulin to activate its receptor and initiate regulation of glucose homeostasis are only partly understood. Here, using ring-closing metathesis, we substitute the A6-A11 disulfide bond of insulin with a rigid, non-reducible dicarba linkage, yielding two distinct stereo-isomers (cis and trans). Remarkably, only the cis isomer displays full insulin potency, rapidly lowering blood glucose in mice (even under insulin-resistant conditions). It also posseses reduced mitogenic activity in vitro. Further biophysical, crystallographic and molecular-dynamics analyses reveal that the A6-A11 bond configuration directly affects the conformational flexibility of insulin A-chain N-terminal helix, dictating insulin's ability to engage its receptor. We reveal that in native insulin, contraction of the C α -C α distance of the flexible A6-A11 cystine allows the A-chain N-terminal helix to unwind to a conformation that allows receptor engagement. This motion is also permitted in the cis isomer, with its shorter C α -C α distance, but prevented in the extended trans analogue. These findings thus illuminate for the first time the allosteric role of the A6-A11 bond in mediating the transition of the hormone to an active conformation, significantly advancing our understanding of insulin action and opening up new avenues for the design of improved therapeutic analogues.
- School of Chemistry, Monash University, Clayton, Victoria, 3800, Australia.
Organizational Affiliation: