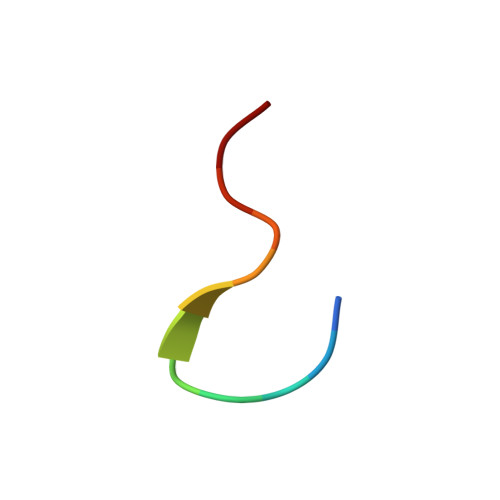
Self-Assembly of Catenanes from Lasso Peptides.
Allen, C.D., Link, A.J.(2016) J Am Chem Soc 138: 14214-14217
- PubMed: 27768305 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/jacs.6b09454
- Primary Citation Related Structures:
5T56 - PubMed Abstract:
Lasso peptides exist naturally in a threaded state as [1]rotaxanes, and we reasoned that lasso peptides cleaved in their loop region could serve as building blocks for catenanes. Mutagenesis of the lasso peptide microcin J25 (MccJ25) with two cysteine residues followed by cleavage of the peptide with trypsin led to a [2]rotaxane structure that self-assembled into a [3]catenane and [4]catenanes at room temperature in aqueous solution. The [3]catenane represents the smallest ring size of a catenane composed solely of polypeptide segments. The NMR structure of the [3]catenane was determined, suggesting that burial of hydrophobic residues may be a driving force for assembly of the catenane structure.
- Departments of Chemical and Biological Engineering and ‡Molecular Biology, Princeton University , Princeton, New Jersey 08544, United States.
Organizational Affiliation: