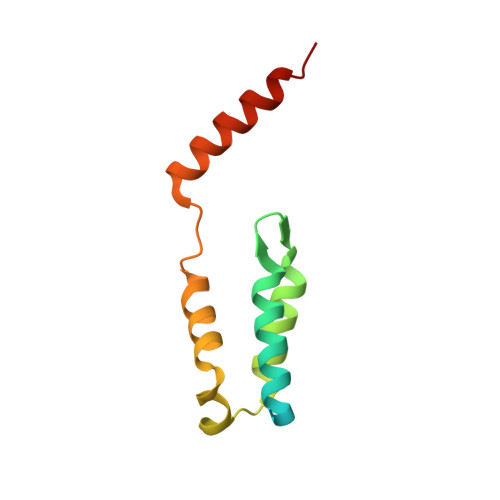
Cloning, purification and structure determination of the HIV integrase-binding domain of lens epithelium-derived growth factor.
Hannon, C., Cruz-Migoni, A., Platonova, O., Owen, R.L., Nettleship, J.E., Miller, A., Carr, S.B., Harris, G., Rabbitts, T.H., Phillips, S.E.V.(2018) Acta Crystallogr F Struct Biol Commun 74: 143-149
- PubMed: 29497017 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X18001553
- Primary Citation Related Structures:
5OYM - PubMed Abstract:
Lens epithelium-derived growth factor (LEDGF)/p75 is the dominant binding partner of HIV-1 integrase in human cells. The crystal structure of the HIV integrase-binding domain (IBD) of LEDGF has been determined in the absence of ligand. IBD was overexpressed in Escherichia coli, purified and crystallized by sitting-drop vapour diffusion. X-ray diffraction data were collected at Diamond Light Source to a resolution of 2.05 Å. The crystals belonged to space group P2 1 , with eight polypeptide chains in the asymmetric unit arranged as an unusual octamer composed of four domain-swapped IBD dimers. IBD exists as a mixture of monomers and dimers in concentrated solutions, but the dimers are unlikely to be biologically relevant.
- Weatherall Institute of Molecular Medicine, University of Oxford, John Radcliffe Hospital, Oxford OX3 9DS, England.
Organizational Affiliation: