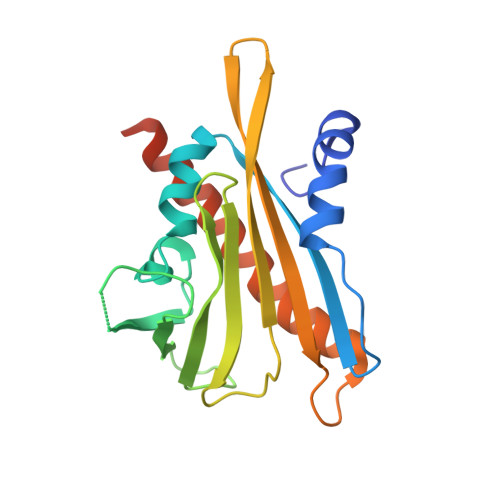
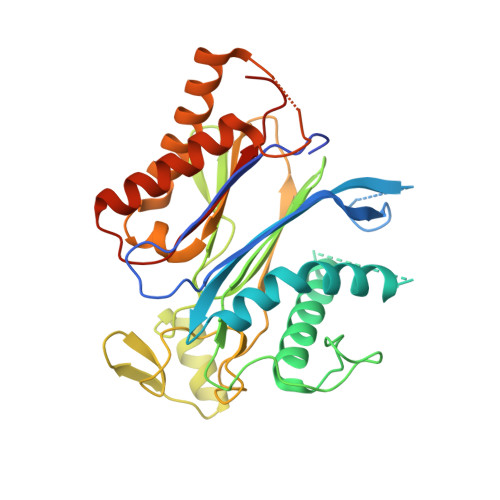
Insights into the in Vitro and in Vivo SAR of Abscisic Acid - Exploring Unprecedented Variations of the Side Chain via Cross-Coupling-Mediated Syntheses
Frackenpohl, J., Grill, E., Bojack, G., Baltz, R., Busch, M., Dittgen, J., Franke, J., Freigang, J., Gonzalez, S., Heinemann, I., Helmke, H., Hills, M., Hohmann, S., von Koskull-Doering, P., Kleemann, J., Lange, G., Lehr, S., Mueller, T., Peschel, E., Poree, F., Schmutzler, D., Schulz, A., Willms, L., Wunschel, C.(2018) European J Org Chem