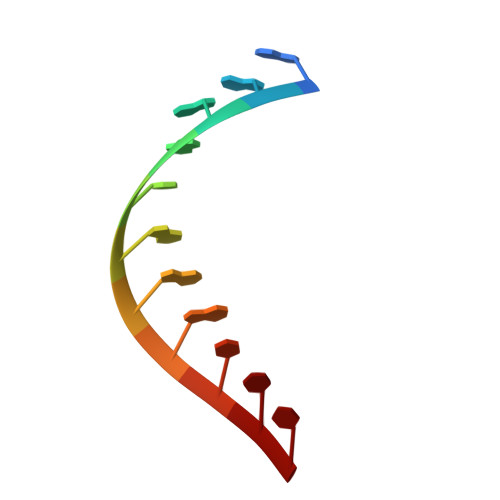
Binding to SMN2 pre-mRNA-protein complex elicits specificity for small molecule splicing modifiers.
Sivaramakrishnan, M., McCarthy, K.D., Campagne, S., Huber, S., Meier, S., Augustin, A., Heckel, T., Meistermann, H., Hug, M.N., Birrer, P., Moursy, A., Khawaja, S., Schmucki, R., Berntenis, N., Giroud, N., Golling, S., Tzouros, M., Banfai, B., Duran-Pacheco, G., Lamerz, J., Hsiu Liu, Y., Luebbers, T., Ratni, H., Ebeling, M., Clery, A., Paushkin, S., Krainer, A.R., Allain, F.H., Metzger, F.(2017) Nat Commun 8: 1476-1476
- PubMed: 29133793 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-017-01559-4
- Primary Citation Related Structures:
5OR0 - PubMed Abstract:
Small molecule splicing modifiers have been previously described that target the general splicing machinery and thus have low specificity for individual genes. Several potent molecules correcting the splicing deficit of the SMN2 (survival of motor neuron 2) gene have been identified and these molecules are moving towards a potential therapy for spinal muscular atrophy (SMA). Here by using a combination of RNA splicing, transcription, and protein chemistry techniques, we show that these molecules directly bind to two distinct sites of the SMN2 pre-mRNA, thereby stabilizing a yet unidentified ribonucleoprotein (RNP) complex that is critical to the specificity of these small molecules for SMN2 over other genes. In addition to the therapeutic potential of these molecules for treatment of SMA, our work has wide-ranging implications in understanding how small molecules can interact with specific quaternary RNA structures.
- F. Hoffmann-La Roche Ltd., Pharma Research & Early Development, Roche Innovation Center Basel, Grenzacherstrasse 124, Basel, 4070, Switzerland.
Organizational Affiliation: