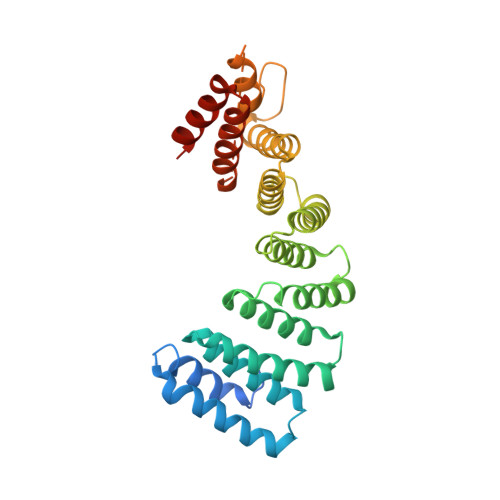
Structural plasticity of the N-terminal capping helix of the TPR domain of kinesin light chain.
Nguyen, T.Q., Chenon, M., Vilela, F., Velours, C., Aumont-Nicaise, M., Andreani, J., Varela, P.F., Llinas, P., Menetrey, J.(2017) PLoS One 12: e0186354-e0186354
- PubMed: 29036226 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0186354
- Primary Citation Related Structures:
5OJ8, 5OJF - PubMed Abstract:
Kinesin1 plays a major role in neuronal transport by recruiting many different cargos through its kinesin light chain (KLC). Various structurally unrelated cargos interact with the conserved tetratricopeptide repeat (TPR) domain of KLC. The N-terminal capping helix of the TPR domain exhibits an atypical sequence and structural features that may contribute to the versatility of the TPR domain to bind different cargos. We determined crystal structures of the TPR domain of both KLC1 and KLC2 encompassing the N-terminal capping helix and show that this helix exhibits two distinct and defined orientations relative to the rest of the TPR domain. Such a difference in orientation gives rise, at the N-terminal part of the groove, to the formation of one hydrophobic pocket, as well as to electrostatic variations at the groove surface. We present a comprehensive structural analysis of available KLC1/2-TPR domain structures that highlights that ligand binding into the groove can be specific of one or the other N-terminal capping helix orientations. Further, structural analysis reveals that the N-terminal capping helix is always involved in crystal packing contacts, especially in a TPR1:TPR1' contact which highlights its propensity to be a protein-protein interaction site. Together, these results underline that the structural plasticity of the N-terminal capping helix might represent a structural determinant for TPR domain structural versatility in cargo binding.
- Laboratoire d'Enzymologie et Biochimie Structurales (LEBS), CNRS, Université Paris-Sud, 1 avenue de la Terrasse, Gif-sur-Yvette, France.
Organizational Affiliation: