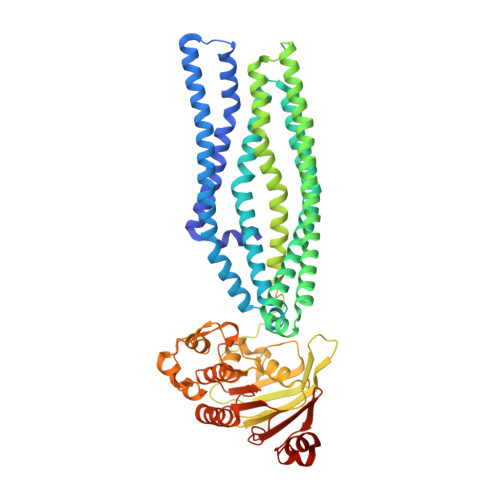
Structural basis for antibacterial peptide self-immunity by the bacterial ABC transporter McjD.
Bountra, K., Hagelueken, G., Choudhury, H.G., Corradi, V., El Omari, K., Wagner, A., Mathavan, I., Zirah, S., Yuan Wahlgren, W., Tieleman, D.P., Schiemann, O., Rebuffat, S., Beis, K.(2017) EMBO J 36: 3062-3079
- PubMed: 28864543 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.15252/embj.201797278
- Primary Citation Related Structures:
5OFP, 5OFR - PubMed Abstract:
Certain pathogenic bacteria produce and release toxic peptides to ensure either nutrient availability or evasion from the immune system. These peptides are also toxic to the producing bacteria that utilize dedicated ABC transporters to provide self-immunity. The ABC transporter McjD exports the antibacterial peptide MccJ25 in Escherichia coli Our previously determined McjD structure provided some mechanistic insights into antibacterial peptide efflux. In this study, we have determined its structure in a novel conformation, apo inward-occluded and a new nucleotide-bound state, high-energy outward-occluded intermediate state, with a defined ligand binding cavity. Predictive cysteine cross-linking in E. coli membranes and PELDOR measurements along the transport cycle indicate that McjD does not undergo major conformational changes as previously proposed for multi-drug ABC exporters. Combined with transport assays and molecular dynamics simulations, we propose a novel mechanism for toxic peptide ABC exporters that only requires the transient opening of the cavity for release of the peptide. We propose that shielding of the cavity ensures that the transporter is available to export the newly synthesized peptides, preventing toxic-level build-up.
- Department of Life Sciences, Imperial College London, London, UK.
Organizational Affiliation: