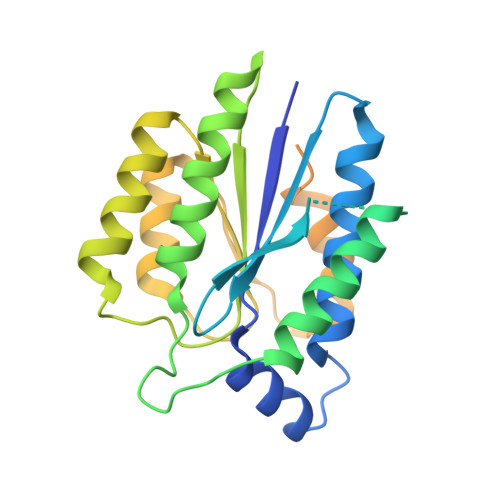
The intricate network between the p34 and p44 subunits is central to the activity of the transcription/DNA repair factor TFIIH.
Radu, L., Schoenwetter, E., Braun, C., Marcoux, J., Koelmel, W., Schmitt, D.R., Kuper, J., Cianferani, S., Egly, J.M., Poterszman, A., Kisker, C.(2017) Nucleic Acids Res 45: 10872-10883
- PubMed: 28977422 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkx743
- Primary Citation Related Structures:
5NUS, 5O85, 5OBZ - PubMed Abstract:
The general transcription factor IIH (TFIIH) is a multi-protein complex and its 10 subunits are engaged in an intricate protein-protein interaction network critical for the regulation of its transcription and DNA repair activities that are so far little understood on a molecular level. In this study, we focused on the p44 and the p34 subunits, which are central for the structural integrity of core-TFIIH. We solved crystal structures of a complex formed by the p34 N-terminal vWA and p44 C-terminal zinc binding domains from Chaetomium thermophilum and from Homo sapiens. Intriguingly, our functional analyses clearly revealed the presence of a second interface located in the C-terminal zinc binding region of p34, which can rescue a disrupted interaction between the p34 vWA and the p44 RING domain. In addition, we demonstrate that the C-terminal zinc binding domain of p34 assumes a central role with respect to the stability and function of TFIIH. Our data reveal a redundant interaction network within core-TFIIH, which may serve to minimize the susceptibility to mutational impairment. This provides first insights why so far no mutations in the p34 or p44 TFIIH-core subunits have been identified that would lead to the hallmark nucleotide excision repair syndromes xeroderma pigmentosum or trichothiodystrophy.
- Institut de Génétique et de Biologie Moléculaire et Cellulaire, UMR 7104 CNRS/Inserm/UdS, BP163, 67404 Illkirch Cedex, C.U. Strasbourg, France.
Organizational Affiliation: