Fructosylation of Hydroxytyrosol by the beta-Fructofuranosidase from Xanthophyllomyces dendrorhous: Insights into the Molecular Basis of the Enzyme Specificity
Ramirez-Escudero, M., Poveda, a., Ballesteros, A.O., Plou, F.J.(2018) ChemCatChem
Experimental Data Snapshot
Starting Model: experimental
View more details
(2018) ChemCatChem
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Beta-fructofuranosidase | 665 | Phaffia rhodozyma | Mutation(s): 1 Gene Names: INV | 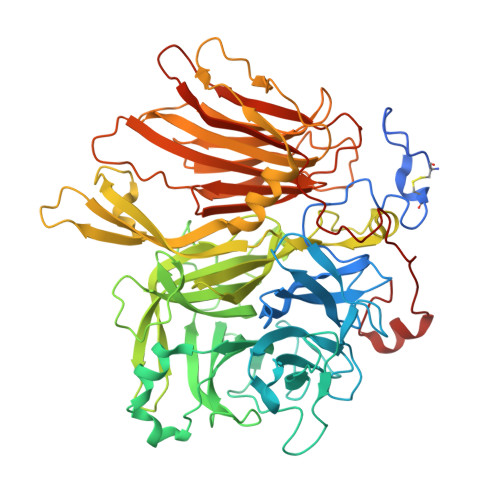 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | J7HDY4 | ||||
Glycosylation | |||||
| Glycosylation Sites: 15 | |||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | 2D Diagram | Glycosylation | D Interactions |
| alpha-D-mannopyranose-(1-3)-alpha-D-mannopyranose-(1-6)-[alpha-D-mannopyranose-(1-3)]beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose | C, F | 6 |  | N-Glycosylation | |
Glycosylation Resources | |||||
GlyTouCan: G09724ZC GlyCosmos: G09724ZC GlyGen: G09724ZC | |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | 2D Diagram | Glycosylation | D Interactions |
| alpha-D-mannopyranose-(1-2)-alpha-D-mannopyranose-(1-2)-alpha-D-mannopyranose-(1-3)-[alpha-D-mannopyranose-(1-2)-alpha-D-mannopyranose-(1-6)-[alpha-D-mannopyranose-(1-3)]alpha-D-mannopyranose-(1-6)]beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose | D, G | 10 |  | N-Glycosylation | |
Glycosylation Resources | |||||
GlyTouCan: G40702WU GlyCosmos: G40702WU GlyGen: G40702WU | |||||
| Ligands 5 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| 9OK Download:Ideal Coordinates CCD File | X [auth A], XA [auth B] | 2-(3,4-dihydroxyphenyl)ethyl beta-D-fructofuranoside C14 H20 O8 LKYHMBCDFKCNAO-YIYPIFLZSA-N |  | ||
| 9WZ Download:Ideal Coordinates CCD File | Y [auth A], YA [auth B] | 2-hydroxy-4-(2-hydroxyethyl)phenyl beta-D-fructofuranoside C14 H20 O8 CNXXGYXVIZNKJW-YIYPIFLZSA-N |  | ||
| NAG Download:Ideal Coordinates CCD File | I [auth A] J [auth A] K [auth A] L [auth A] LA [auth B] | 2-acetamido-2-deoxy-beta-D-glucopyranose C8 H15 N O6 OVRNDRQMDRJTHS-FMDGEEDCSA-N |  | ||
| 975 Download:Ideal Coordinates CCD File | Z [auth A], ZA [auth B] | 4-(2-hydroxyethyl)benzene-1,2-diol C8 H10 O3 JUUBCHWRXWPFFH-UHFFFAOYSA-N |  | ||
| EDO Download:Ideal Coordinates CCD File | AA [auth A] AB [auth B] BA [auth A] BB [auth B] CA [auth A] | 1,2-ETHANEDIOL C2 H6 O2 LYCAIKOWRPUZTN-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 74.616 | α = 90 |
| b = 205.356 | β = 90 |
| c = 146.94 | γ = 90 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| XDS | data reduction |
| Aimless | data scaling |