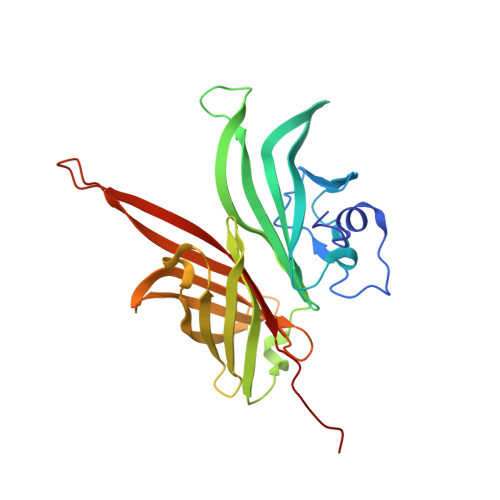
Structure-based design of chimeric antigens for multivalent protein vaccines.
Hollingshead, S., Jongerius, I., Exley, R.M., Johnson, S., Lea, S.M., Tang, C.M.(2018) Nat Commun 9: 1051-1051
- PubMed: 29535307 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-018-03146-7
- Primary Citation Related Structures:
5NQP, 5NQX, 5NQY, 5NQZ - PubMed Abstract:
There is an urgent need to develop vaccines against pathogenic bacteria. However, this is often hindered by antigenic diversity and difficulties encountered manufacturing membrane proteins. Here we show how to use structure-based design to develop chimeric antigens (ChAs) for subunit vaccines. ChAs are generated against serogroup B Neisseria meningitidis (MenB), the predominant cause of meningococcal disease in wealthy countries. MenB ChAs exploit factor H binding protein (fHbp) as a molecular scaffold to display the immunogenic VR2 epitope from the integral membrane protein PorA. Structural analyses demonstrate fHbp is correctly folded and the PorA VR2 epitope adopts an immunogenic conformation. In mice, immunisation with ChAs generates fHbp and PorA antibodies that recognise the antigens expressed by clinical MenB isolates; these antibody responses correlate with protection against meningococcal disease. Application of ChAs is therefore a potentially powerful approach to develop multivalent subunit vaccines, which can be tailored to circumvent pathogen diversity.
- Sir William Dunn School of Pathology, University of Oxford, South Parks Road, Oxford, OX1 3RE, UK.
Organizational Affiliation: