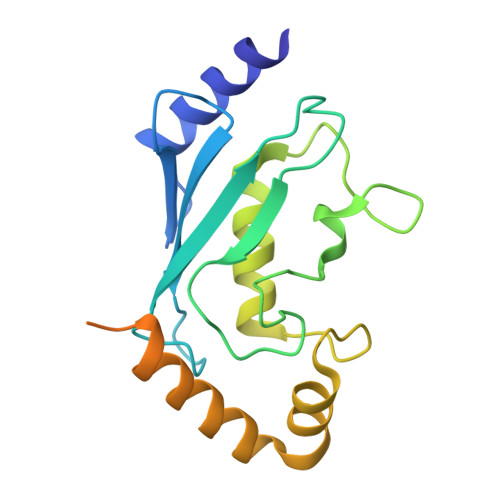
Allosteric Targeting of the Fanconi Anemia Ubiquitin-Conjugating Enzyme Ube2T by Fragment Screening.
Morreale, F.E., Bortoluzzi, A., Chaugule, V.K., Arkinson, C., Walden, H., Ciulli, A.(2017) J Med Chem 60: 4093-4098
- PubMed: 28437106 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.jmedchem.7b00147
- Primary Citation Related Structures:
5NGZ - PubMed Abstract:
Ube2T is the E2 ubiquitin-conjugating enzyme of the Fanconi anemia DNA repair pathway and it is overexpressed in several cancers, representing an attractive target for the development of inhibitors. Despite the extensive efforts in targeting the ubiquitin system, very few E2 binders have currently been discovered. Herein we report the identification of a new allosteric pocket on Ube2T through a fragment screening using biophysical methods. Several fragments binding to this site inhibit ubiquitin conjugation in vitro.
- MRC Protein Phosphorylation and Ubiquitylation Unit, School of Life Sciences, University of Dundee , Dundee DD1 5EH, United Kingdom.
Organizational Affiliation: