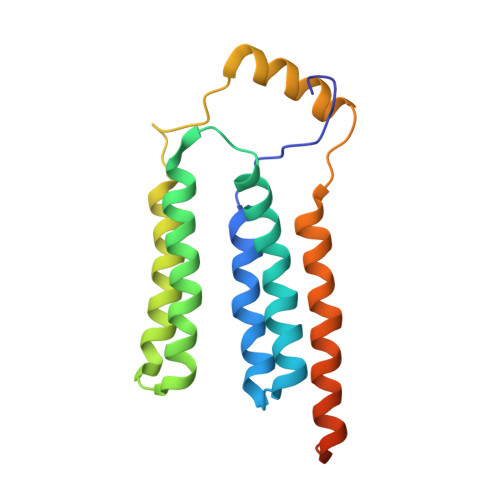
Solution NMR structure of yeast Rcf1, a protein involved in respiratory supercomplex formation.
Zhou, S., Pettersson, P., Huang, J., Sjoholm, J., Sjostrand, D., Pomes, R., Hogbom, M., Brzezinski, P., Maler, L., Adelroth, P.(2018) Proc Natl Acad Sci U S A 115: 3048-3053
- PubMed: 29507228 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1712061115
- Primary Citation Related Structures:
5NF8 - PubMed Abstract:
The Saccharomyces cerevisiae respiratory supercomplex factor 1 (Rcf1) protein is located in the mitochondrial inner membrane where it is involved in formation of supercomplexes composed of respiratory complexes III and IV. We report the solution structure of Rcf1, which forms a dimer in dodecylphosphocholine (DPC) micelles, where each monomer consists of a bundle of five transmembrane (TM) helices and a short flexible soluble helix (SH). Three TM helices are unusually charged and provide the dimerization interface consisting of 10 putative salt bridges, defining a "charge zipper" motif. The dimer structure is supported by molecular dynamics (MD) simulations in DPC, although the simulations show a more dynamic dimer interface than the NMR data. Furthermore, CD and NMR data indicate that Rcf1 undergoes a structural change when reconstituted in liposomes, which is supported by MD data, suggesting that the dimer structure is unstable in a planar membrane environment. Collectively, these data indicate a dynamic monomer-dimer equilibrium. Furthermore, the Rcf1 dimer interacts with cytochrome c , suggesting a role as an electron-transfer bridge between complexes III and IV. The Rcf1 structure will help in understanding its functional roles at a molecular level.
- Department of Biochemistry and Biophysics, Stockholm University, 10691 Stockholm, Sweden.
Organizational Affiliation: