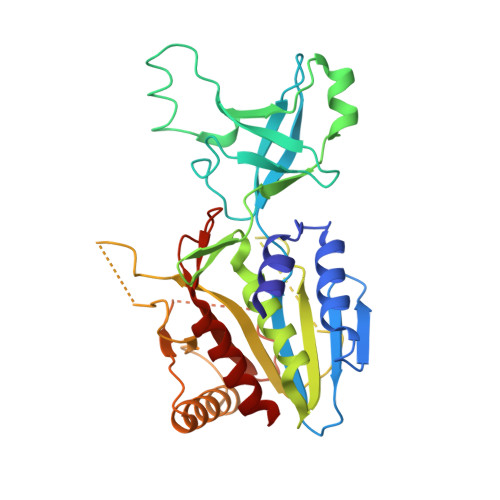
X-Ray Crystallography to Study the Oligomeric State Transition of the Thermotoga maritima M42 Aminopeptidase TmPep1050.
Dutoit, R., Brandt, N., Van Elder, D., Droogmans, L.(2020) J Vis Exp
- PubMed: 32478746 Search on PubMed
- DOI: https://doi.org/10.3791/61288
- Primary Citation Related Structures:
5NE9 - PubMed Abstract:
The M42 aminopeptidases form functionally active complexes made of 12 subunits. Their assembly process appears to be regulated by their metal ion cofactors triggering a dimer-dodecamer transition. Upon metal ion binding, several structural modifications occur in the active site and at the interaction interface, shaping dimers to promote the self-assembly. To observe such modifications, stable oligomers must be isolated prior to structural study. Reported here is a method that allows the purification of stable dodecamers and dimers of TmPep1050, an M42 aminopeptidase of T. maritima, and their structure determination by X-ray crystallography. Dimers were prepared from dodecamers by removing metal ions with a chelating agent. Without their cofactor, dodecamers became less stable and were fully dissociated upon heating. The oligomeric structures were solved by the straightforward molecular replacement approach. To illustrate the methodology, the structure of a TmPep1050 variant, totally impaired in metal ion binding, is presented showing no further breakdown of dimers to monomers.
- Laboratory of Microbiology, Department of Molecular Biology, Université Libre de Bruxelles; Labiris Institut de Recherche; Raphael.Dutoit@ulb.ac.be.
Organizational Affiliation: