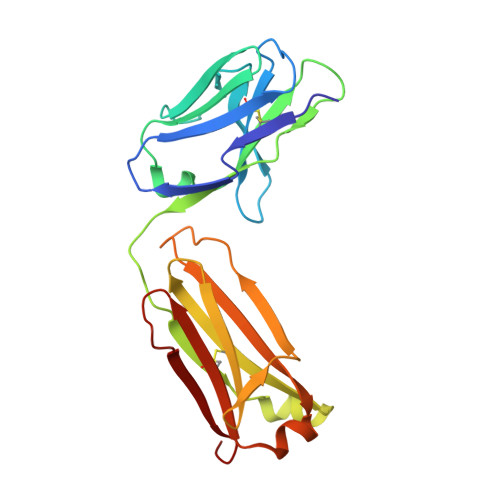
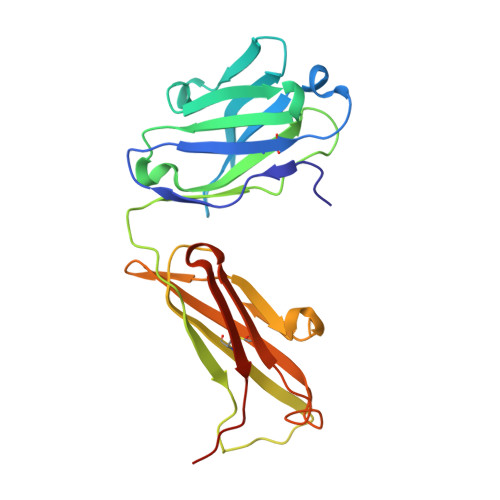
Tight Molecular Recognition of Benzo[a]pyrene by a High-Affinity Antibody.
Eichinger, A., Neumaier, I., Pschenitza, M., Niessner, R., Knopp, D., Skerra, A.(2017) Angew Chem Int Ed Engl 56: 10592-10597
- PubMed: 28603847 Search on PubMed
- DOI: https://doi.org/10.1002/anie.201703893
- Primary Citation Related Structures:
5NBW - PubMed Abstract:
Benzo[a]pyrene, which is produced during the incomplete combustion of organic material, is an abundant noxious pollutant because of its carcinogenic metabolic degradation products. The high-affinity (K D ≈3 nm) monoclonal antibody 22F12 allows facile bioanalytical quantification of benzo[a]pyrene even in complex matrices. We report the functional and X-ray crystallographic analysis of 22F12 in complex with 3-hydroxybenzo[a]pyrene after cloning of the V-genes and production as a recombinant Fab fragment. The polycyclic aromatic hydrocarbon is bound in a deep pocket between the light and heavy chains, surrounded mainly by aromatic and aliphatic amino acid side chains. Interestingly, the hapten-antibody interface is less densely packed than expected and reveals polar, H-bond-like interactions with the polycyclic aromatic π-electron system, which may allow the antibody to maintain a large, predominantly hydrophobic binding site in an aqueous environment while providing sufficient complementarity to its ligand.
- Munich Center for Integrated Protein Science, CIPS-M, und Lehrstuhl für Biologische Chemie, Technische Universität München, 85354, Freising, Weihenstephan, Germany.
Organizational Affiliation: