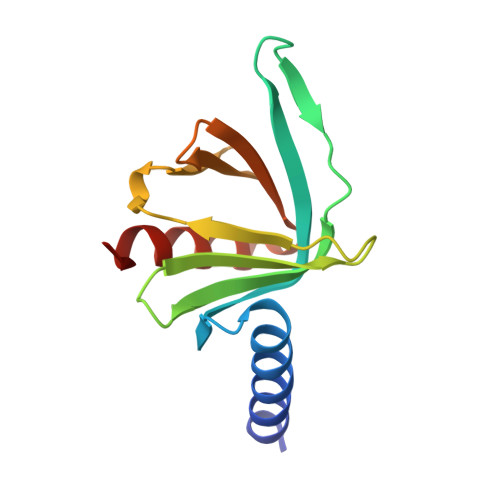
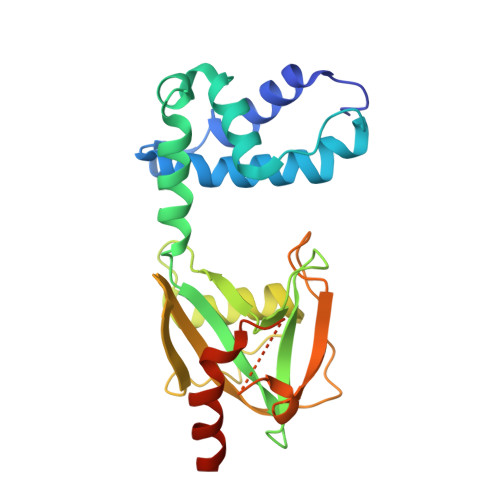
Changes in conformational equilibria regulate the activity of the Dcp2 decapping enzyme.
Wurm, J.P., Holdermann, I., Overbeck, J.H., Mayer, P.H.O., Sprangers, R.(2017) Proc Natl Acad Sci U S A 114: 6034-6039
- PubMed: 28533364 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1704496114
- Primary Citation Related Structures:
5N2V - PubMed Abstract:
Crystal structures of enzymes are indispensable to understanding their mechanisms on a molecular level. It, however, remains challenging to determine which structures are adopted in solution, especially for dynamic complexes. Here, we study the bilobed decapping enzyme Dcp2 that removes the 5' cap structure from eukaryotic mRNA and thereby efficiently terminates gene expression. The numerous Dcp2 structures can be grouped into six states where the domain orientation between the catalytic and regulatory domains significantly differs. Despite this wealth of structural information it is not possible to correlate these states with the catalytic cycle or the activity of the enzyme. Using methyl transverse relaxation-optimized NMR spectroscopy, we demonstrate that only three of the six domain orientations are present in solution, where Dcp2 adopts an open, a closed, or a catalytically active state. We show how mRNA substrate and the activator proteins Dcp1 and Edc1 influence the dynamic equilibria between these states and how this modulates catalytic activity. Importantly, the active state of the complex is only stably formed in the presence of both activators and the mRNA substrate or the m7GDP decapping product, which we rationalize based on a crystal structure of the Dcp1:Dcp2:Edc1:m7GDP complex. Interestingly, we find that the activating mechanisms in Dcp2 also result in a shift of the substrate specificity from bacterial to eukaryotic mRNA.
- Max Planck Institute for Developmental Biology, 72076 Tuebingen, Germany.
Organizational Affiliation: