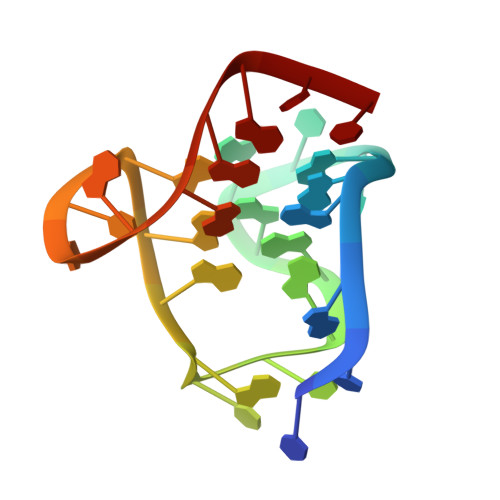
Solution NMR Structure of a Ligand/Hybrid-2-G-Quadruplex Complex Reveals Rearrangements that Affect Ligand Binding.
Wirmer-Bartoschek, J., Bendel, L.E., Jonker, H.R.A., Grun, J.T., Papi, F., Bazzicalupi, C., Messori, L., Gratteri, P., Schwalbe, H.(2017) Angew Chem Int Ed Engl 56: 7102-7106
- PubMed: 28524432 Search on PubMed
- DOI: https://doi.org/10.1002/anie.201702135
- Primary Citation Related Structures:
5MVB - PubMed Abstract:
Telomeric G-quadruplexes have recently emerged as drug targets in cancer research. Herein, we present the first NMR structure of a telomeric DNA G-quadruplex that adopts the biologically relevant hybrid-2 conformation in a ligand-bound state. We solved the complex with a metalorganic gold(III) ligand that stabilizes G-quadruplexes. Analysis of the free and bound structures reveals structural changes in the capping region of the G-quadruplex. The ligand is sandwiched between one terminal G-tetrad and a flanking nucleotide. This complex structure involves a major structural rearrangement compared to the free G-quadruplex structure as observed for other G-quadruplexes in different conformations, invalidating simple docking approaches to ligand-G-quadruplex structure determination.
- Institute for Organic Chemistry and Chemical Biology, Center of Biomolecular Magnetic Resonance (BMRZ), Goethe University Frankfurt/Main, Max-von-Laue-Strasse 7, 60439, Frankfurt, Germany.
Organizational Affiliation: