Discovery of 2-[[2-Ethyl-6-[4-[2-(3-hydroxyazetidin-1-yl)-2-oxoethyl]piperazin-1-yl]-8-methylimidazo[1,2-a]pyridin-3-yl]methylamino]-4-(4-fluorophenyl)thiazole-5-carbonitrile (GLPG1690), a First-in-Class Autotaxin Inhibitor Undergoing Clinical Evaluation for the Treatment of Idiopathic Pulmonary Fibrosis.
Desroy, N., Housseman, C., Bock, X., Joncour, A., Bienvenu, N., Cherel, L., Labeguere, V., Rondet, E., Peixoto, C., Grassot, J.M., Picolet, O., Annoot, D., Triballeau, N., Monjardet, A., Wakselman, E., Roncoroni, V., Le Tallec, S., Blanque, R., Cottereaux, C., Vandervoort, N., Christophe, T., Mollat, P., Lamers, M., Auberval, M., Hrvacic, B., Ralic, J., Oste, L., van der Aar, E., Brys, R., Heckmann, B.(2017) J Med Chem 60: 3580-3590
- PubMed: 28414242 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.7b00032
- Primary Citation Related Structures:
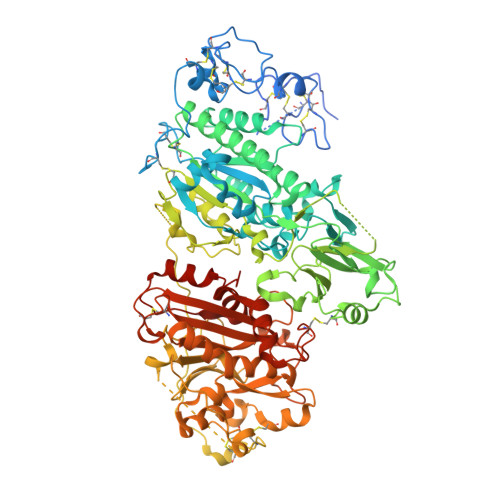
5MHP - PubMed Abstract:
Autotaxin is a circulating enzyme with a major role in the production of lysophosphatic acid (LPA) species in blood. A role for the autotaxin/LPA axis has been suggested in many disease areas including pulmonary fibrosis. Structural modifications of the known autotaxin inhibitor lead compound 1, to attenuate hERG inhibition, remove CYP3A4 time-dependent inhibition, and improve pharmacokinetic properties, led to the identification of clinical candidate GLPG1690 (11). Compound 11 was able to cause a sustained reduction of LPA levels in plasma in vivo and was shown to be efficacious in a bleomycin-induced pulmonary fibrosis model in mice and in reducing extracellular matrix deposition in the lung while also reducing LPA 18:2 content in bronchoalveolar lavage fluid. Compound 11 is currently being evaluated in an exploratory phase 2a study in idiopathic pulmonary fibrosis patients.
- Galapagos SASU , 102 Avenue Gaston Roussel, 93230 Romainville, France.
Organizational Affiliation: