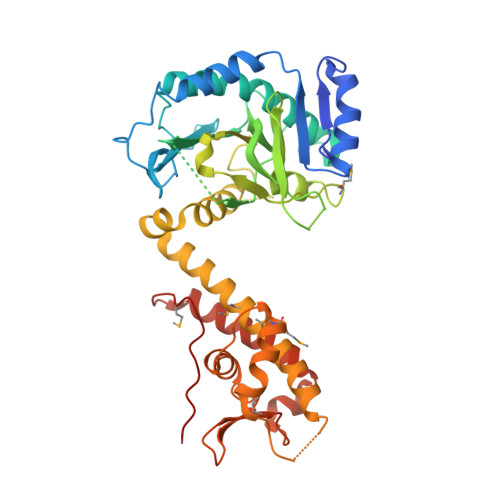
Cdt1 stabilizes an open MCM ring for helicase loading.
Frigola, J., He, J., Kinkelin, K., Pye, V.E., Renault, L., Douglas, M.E., Remus, D., Cherepanov, P., Costa, A., Diffley, J.F.X.(2017) Nat Commun 8: 15720-15720
- PubMed: 28643783 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms15720
- Primary Citation Related Structures:
5ME9, 5MEA, 5MEB, 5MEC - PubMed Abstract:
ORC, Cdc6 and Cdt1 act together to load hexameric MCM, the motor of the eukaryotic replicative helicase, into double hexamers at replication origins. Here we show that Cdt1 interacts with MCM subunits Mcm2, 4 and 6, which both destabilizes the Mcm2-5 interface and inhibits MCM ATPase activity. Using X-ray crystallography, we show that Cdt1 contains two winged-helix domains in the C-terminal half of the protein and a catalytically inactive dioxygenase-related N-terminal domain, which is important for MCM loading, but not for subsequent replication. We used these structures together with single-particle electron microscopy to generate three-dimensional models of MCM complexes. These show that Cdt1 stabilizes MCM in a left-handed spiral open at the Mcm2-5 gate. We propose that Cdt1 acts as a brace, holding MCM open for DNA entry and bound to ATP until ORC-Cdc6 triggers ATP hydrolysis by MCM, promoting both Cdt1 ejection and MCM ring closure.
- Chromosome Replication Laboratory, The Francis Crick Institute, 1 Midland Road, London NW1 1AT, UK.
Organizational Affiliation: