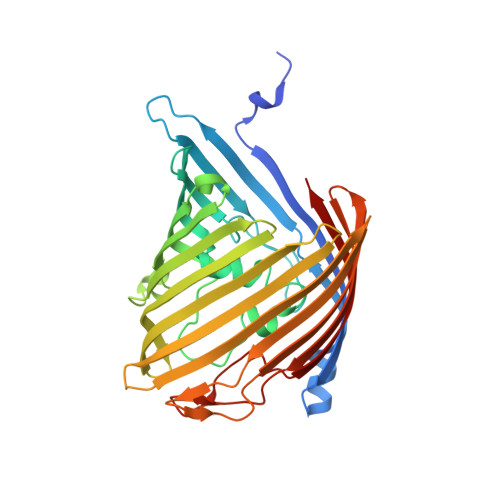
Structural basis for chitin acquisition by marine Vibrio species.
Aunkham, A., Zahn, M., Kesireddy, A., Pothula, K.R., Schulte, A., Basle, A., Kleinekathofer, U., Suginta, W., van den Berg, B.(2018) Nat Commun 9: 220-220
- PubMed: 29335469 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-017-02523-y
- Primary Citation Related Structures:
5MDO, 5MDP, 5MDQ, 5MDR, 5MDS - PubMed Abstract:
Chitin, an insoluble polymer of N-acetylglucosamine, is one of the most abundant biopolymers on Earth. By degrading chitin, chitinolytic bacteria such as Vibrio harveyi are critical for chitin recycling and maintenance of carbon and nitrogen cycles in the world's oceans. A decisive step in chitin degradation is the uptake of chito-oligosaccharides by an outer membrane protein channel named chitoporin (ChiP). Here, we report X-ray crystal structures of ChiP from V. harveyi in the presence and absence of chito-oligosaccharides. Structures without bound sugar reveal a trimeric assembly with an unprecedented closing of the transport pore by the N-terminus of a neighboring subunit. Substrate binding ejects the pore plug to open the transport channel. Together with molecular dynamics simulations, electrophysiology and in vitro transport assays our data provide an explanation for the exceptional affinity of ChiP for chito-oligosaccharides and point to an important role of the N-terminal gate in substrate transport.
- Biochemistry-Electrochemistry Research Unit, Institute of Science, Suranaree University of Technology, Nakhon Ratchasima, 30000, Thailand.
Organizational Affiliation: