Bivalent Llama Single-Domain Antibody Fragments against Tumor Necrosis Factor Have Picomolar Potencies due to Intramolecular Interactions.
Beirnaert, E., Desmyter, A., Spinelli, S., Lauwereys, M., Aarden, L., Dreier, T., Loris, R., Silence, K., Pollet, C., Cambillau, C., de Haard, H.(2017) Front Immunol 8: 867-867
- PubMed: 28824615 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3389/fimmu.2017.00867
- Primary Citation Related Structures:
5M2I, 5M2J, 5M2M - PubMed Abstract:
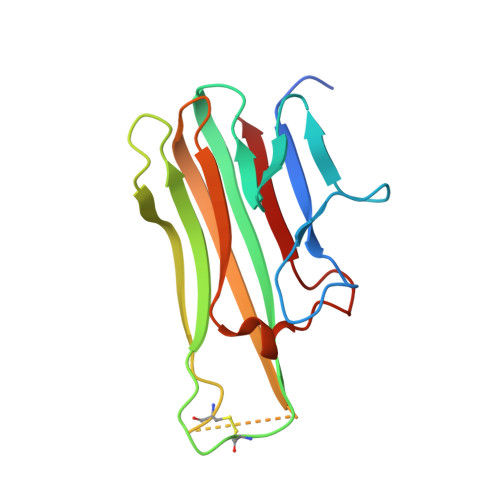
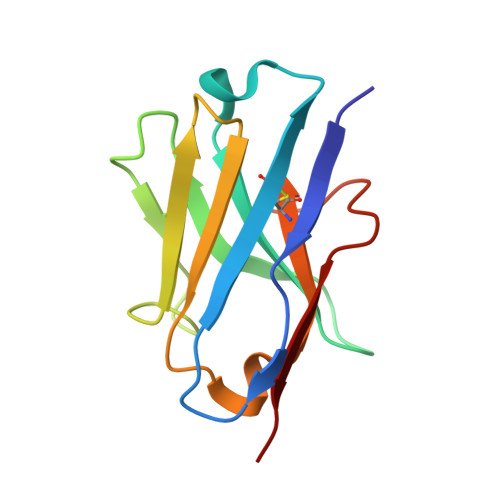
The activity of tumor necrosis factor (TNF), a cytokine involved in inflammatory pathologies, can be inhibited by antibodies or trap molecules. Herein, llama-derived variable heavy-chain domains of heavy-chain antibody (VHH, also called Nanobodies™) were generated for the engineering of bivalent constructs, which antagonize the binding of TNF to its receptors with picomolar potencies. Three monomeric VHHs (VHH#1, VHH#2, and VHH#3) were characterized in detail and found to bind TNF with sub-nanomolar affinities. The crystal structures of the TNF-VHH complexes demonstrate that VHH#1 and VHH#2 share the same epitope, at the center of the interaction area of TNF with its TNFRs, while VHH#3 binds to a different, but partially overlapping epitope. These structures rationalize our results obtained with bivalent constructs in which two VHHs were coupled via linkers of different lengths. Contrary to conventional antibodies, these bivalent Nanobody™ constructs can bind to a single trimeric TNF, thus binding with avidity and blocking two of the three receptor binding sites in the cytokine. The different mode of binding to antigen and the engineering into bivalent constructs supports the design of highly potent VHH-based therapeutic entities.
- Ablynx NV, Ghent, Belgium.
Organizational Affiliation: