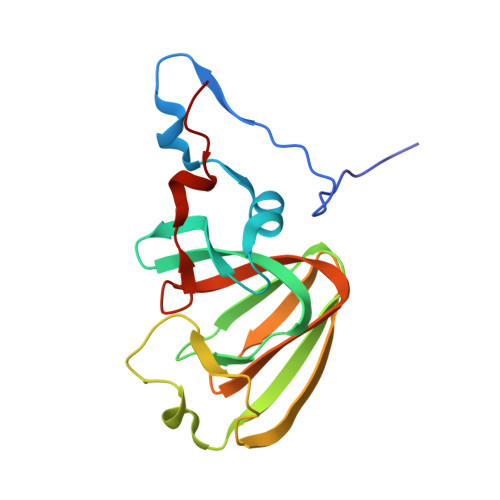
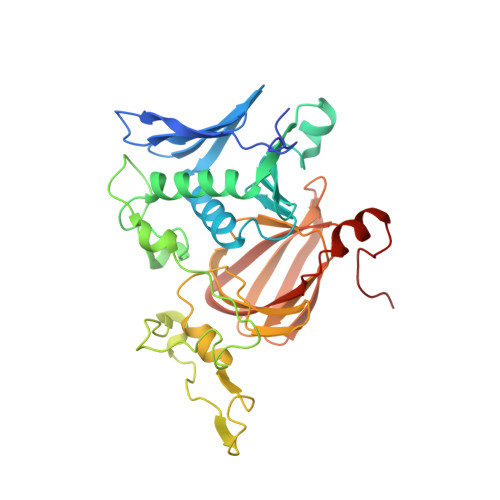
The crystal structures of native hydroquinone 1,2-dioxygenase from Sphingomonas sp. TTNP3 and of substrate and inhibitor complexes.
Ferraroni, M., Da Vela, S., Kolvenbach, B.A., Corvini, P.F., Scozzafava, A.(2017) Biochim Biophys Acta 1865: 520-530
- PubMed: 28232026 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbapap.2017.02.013
- Primary Citation Related Structures:
5M21, 5M22, 5M26, 5M4O - PubMed Abstract:
The crystal structure of hydroquinone 1,2-dioxygenase, a Fe(II) ring cleaving dioxygenase from Sphingomonas sp. strain TTNP3, which oxidizes a wide range of hydroquinones to the corresponding 4-hydroxymuconic semialdehydes, has been solved by Molecular Replacement, using the coordinates of PnpCD from Pseudomonas sp. strain WBC-3. The enzyme is a heterotetramer, constituted of two subunits α and two β of 19 and 38kDa, respectively. Both the two subunits fold as a cupin, but that of the small α subunit lacks a competent metal binding pocket. Two tetramers are present in the asymmetric unit. Each of the four β subunits in the asymmetric unit binds one Fe(II) ion. The iron ion in each β subunit is coordinated to three protein residues, His258, Glu264, and His305 and a water molecule. The crystal structures of the complexes with the substrate methylhydroquinone, obtained under anaerobic conditions, and with the inhibitors 4-hydroxybenzoate and 4-nitrophenol were also solved. The structures of the native enzyme and of the complexes present significant differences in the active site region compared to PnpCD, the other hydroquinone 1,2-dioxygenase of known structure, and in particular they show a different coordination at the metal center.
- Dipartimento di Chimica "Ugo Schiff", Università di Firenze, Via della Lastruccia 3, I-50019, Sesto Fiorentino, FI, Italy. Electronic address: marta.ferraroni@unifi.it.
Organizational Affiliation: