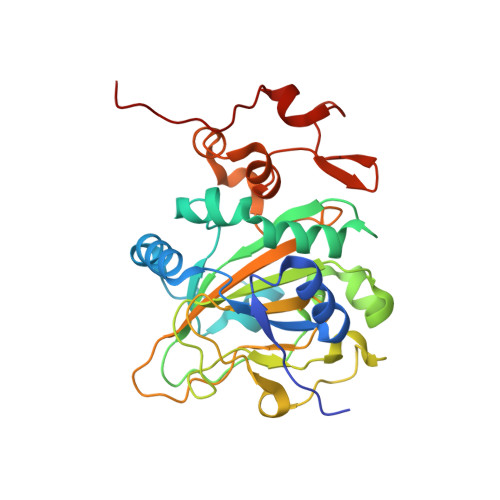
Structural characterization of EasH (Aspergillus japonicus) - an oxidase involved in cycloclavine biosynthesis.
Jakubczyk, D., Caputi, L., Stevenson, C.E., Lawson, D.M., O'Connor, S.E.(2016) Chem Commun (Camb) 52: 14306-14309
- PubMed: 27885368 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1039/c6cc08438a
- Primary Citation Related Structures:
5M0T - PubMed Abstract:
Aj_EasH is a non-heme iron- and α-keto-glutarate-dependent oxidase that is responsible for an unusual cyclopropyl ring formation in the biosynthesis of the fungal ergot alkaloid cycloclavine. The three dimensional structure of Aj_EasH (2.2 Å resolution) reported here provides insight into the mechanism of this unusual and complex reaction.
- Department of Biological Chemistry, John Innes Centre, Norwich Research Park, Norwich NR4 7UH, UK. Sarah.oconnor@jic.ac.uk.
Organizational Affiliation: