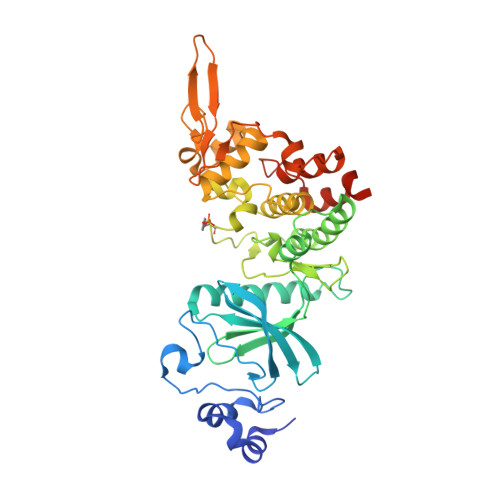
An Unusual Binding Model of the Methyl 9-Anilinothiazolo[5,4-f] quinazoline-2-carbimidates (EHT 1610 and EHT 5372) Confers High Selectivity for Dual-Specificity Tyrosine Phosphorylation-Regulated Kinases.
Chaikuad, A., Diharce, J., Schroder, M., Foucourt, A., Leblond, B., Casagrande, A.S., Desire, L., Bonnet, P., Knapp, S., Besson, T.(2016) J Med Chem 59: 10315-10321
- PubMed: 27766861 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.6b01083
- Primary Citation Related Structures:
5LXC, 5LXD - PubMed Abstract:
Methyl 9-anilinothiazolo[5,4-f]quinazoline-2-carbimidates 1 (EHT 5372) and 2 (EHT 1610) are strong inhibitors of DYRK's family kinases. The crystal structures of the complex revealed a noncanonical binding mode of compounds 1 and 2 in DYRK2, explaining the remarkable selectivity and potency of these inhibitors. The structural data and comparison presented here provide therefore a template for further improvement of this inhibitor class and for the development of novel inhibitors selectively targeting DYRK kinases.
- Target Discovery Institute (TDI), and Structural Genomics Consortium (SGC), University of Oxford , Old Road Campus Research Building, Oxford OX3 7DQ, U.K.
Organizational Affiliation: