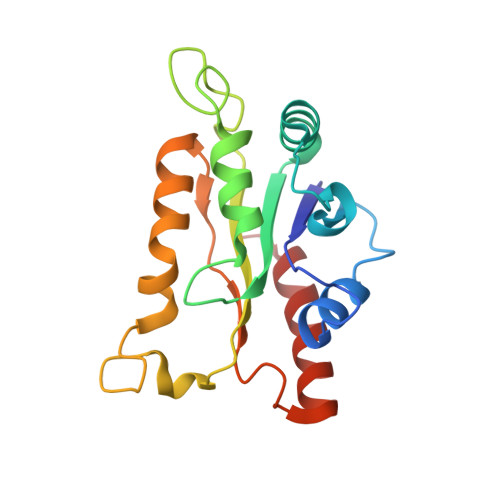
Structure and biocatalytic scope of thermophilic flavin-dependent halogenase and flavin reductase enzymes.
Menon, B.R., Latham, J., Dunstan, M.S., Brandenburger, E., Klemstein, U., Leys, D., Karthikeyan, C., Greaney, M.F., Shepherd, S.A., Micklefield, J.(2016) Org Biomol Chem 14: 9354-9361
- PubMed: 27714222 Search on PubMed
- DOI: https://doi.org/10.1039/c6ob01861k
- Primary Citation Related Structures:
5LV9, 5LVA - PubMed Abstract:
Flavin-dependent halogenase (Fl-Hal) enzymes have been shown to halogenate a range of synthetic as well as natural aromatic compounds. The exquisite regioselectively of Fl-Hal enzymes can provide halogenated building blocks which are inaccessible using standard halogenation chemistries. Consequently, Fl-Hal are potentially useful biocatalysts for the chemoenzymatic synthesis of pharmaceuticals and other valuable products, which are derived from haloaromatic precursors. However, the application of Fl-Hal enzymes, in vitro, has been hampered by their poor catalytic activity and lack of stability. To overcome these issues, we identified a thermophilic tryptophan halogenase (Th-Hal), which has significantly improved catalytic activity and stability, compared with other Fl-Hal characterised to date. When used in combination with a thermostable flavin reductase, Th-Hal can efficiently halogenate a number of aromatic substrates. X-ray crystal structures of Th-Hal, and the reductase partner (Th-Fre), provide insights into the factors that contribute to enzyme stability, which could guide the discovery and engineering of more robust and productive halogenase biocatalysts.
- School of Chemistry and Manchester Institute of Biotechnology, The University of Manchester, 131 Princess Street, Manchester, M1 7DN, UK. jason.micklefield@manchester.ac.uk.
Organizational Affiliation: