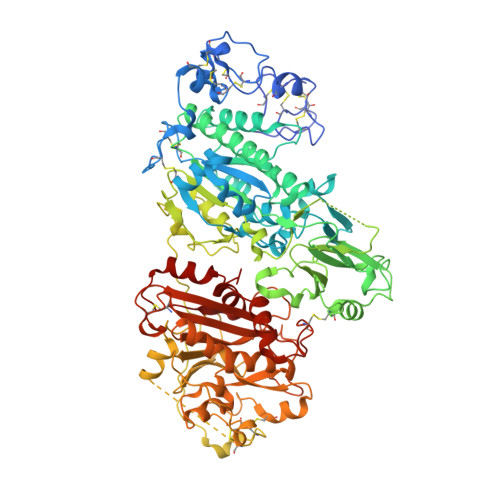
Structure-Activity Relationships of Small Molecule Autotaxin Inhibitors with a Discrete Binding Mode.
Miller, L.M., Keune, W.J., Castagna, D., Young, L.C., Duffy, E.L., Potjewyd, F., Salgado-Polo, F., Engel Garcia, P., Semaan, D., Pritchard, J.M., Perrakis, A., Macdonald, S.J., Jamieson, C., Watson, A.J.(2017) J Med Chem 60: 722-748
- PubMed: 27982588 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.6b01597
- Primary Citation Related Structures:
5LQQ - PubMed Abstract:
Autotaxin (ATX) is a secreted enzyme responsible for the hydrolysis of lysophosphatidylcholine (LPC) to the bioactive lysophosphatidic acid (LPA) and choline. The ATX-LPA signaling pathway is implicated in cell survival, migration, and proliferation; thus, the inhibition of ATX is a recognized therapeutic target for a number of diseases including fibrotic diseases, cancer, and inflammation, among others. Many of the developed synthetic inhibitors for ATX have resembled the lipid chemotype of the native ligand; however, a small number of inhibitors have been described that deviate from this common scaffold. Herein, we report the structure-activity relationships (SAR) of a previously reported small molecule ATX inhibitor. We show through enzyme kinetics studies that analogues of this chemotype are noncompetitive inhibitors, and by using a crystal structure with ATX we confirm the discrete binding mode.
- WestCHEM, Department of Pure and Applied Chemistry, University of Strathclyde , Thomas Graham Building, 295 Cathedral Street, Glasgow G1 1XL, U.K.
Organizational Affiliation: