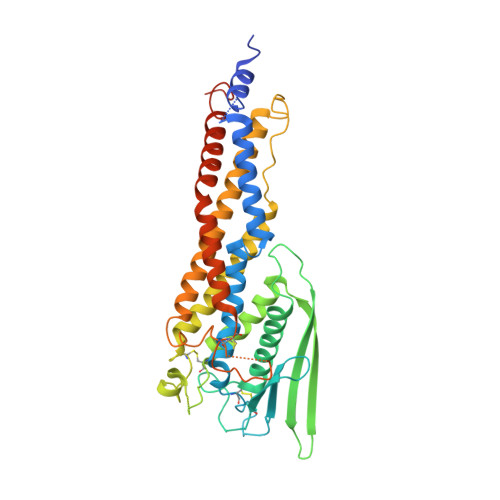
Helicobacter pylori adhesin HopQ engages in a virulence-enhancing interaction with human CEACAMs.
Javaheri, A., Kruse, T., Moonens, K., Mejias-Luque, R., Debraekeleer, A., Asche, C.I., Tegtmeyer, N., Kalali, B., Bach, N.C., Sieber, S.A., Hill, D.J., Koniger, V., Hauck, C.R., Moskalenko, R., Haas, R., Busch, D.H., Klaile, E., Slevogt, H., Schmidt, A., Backert, S., Remaut, H., Singer, B.B., Gerhard, M.(2016) Nat Microbiol 2: 16189-16189
- PubMed: 27748768 Search on PubMed
- DOI: https://doi.org/10.1038/nmicrobiol.2016.189
- Primary Citation Related Structures:
5LP2 - PubMed Abstract:
Helicobacter pylori specifically colonizes the human gastric epithelium and is the major causative agent for ulcer disease and gastric cancer development. Here, we identify members of the carcinoembryonic antigen-related cell adhesion molecule (CEACAM) family as receptors of H. pylori and show that HopQ is the surface-exposed adhesin that specifically binds human CEACAM1, CEACAM3, CEACAM5 and CEACAM6. HopQ-CEACAM binding is glycan-independent and targeted to the N-domain. H. pylori binding induces CEACAM1-mediated signalling, and the HopQ-CEACAM1 interaction enables translocation of the virulence factor CagA into host cells and enhances the release of pro-inflammatory mediators such as interleukin-8. Based on the crystal structure of HopQ, we found that a β-hairpin insertion (HopQ-ID) in HopQ's extracellular 3+4 helix bundle domain is important for CEACAM binding. A peptide derived from this domain competitively inhibits HopQ-mediated activation of the Cag virulence pathway, as genetic or antibody-mediated abrogation of the HopQ function shows. Together, our data suggest the HopQ-CEACAM1 interaction to be a potentially promising novel therapeutic target to combat H. pylori-associated diseases.
- Institute for Medical Microbiology, Immunology and Hygiene, Technische Universität München, 81675 Munich, Germany.
Organizational Affiliation: