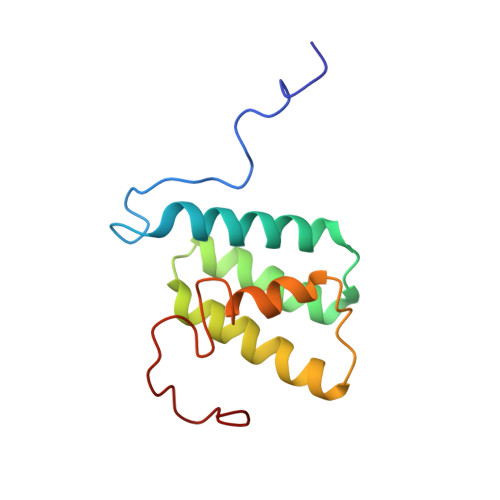
Nuclear Magnetic Resonance Structure and Mutational Analysis of the Lactococcin A Immunity Protein.
Kristiansen, P.E., Persson, C., Fuochi, V., Pedersen, A., Karlsson, G.B., Nissen-Meyer, J., Oppegard, C.(2016) Biochemistry 55: 6250-6257
- PubMed: 27808503 Search on PubMed
- DOI: https://doi.org/10.1021/acs.biochem.6b00848
- Primary Citation Related Structures:
5LFI - PubMed Abstract:
The class IId bacteriocin lactococcin A and the pediocin-like bacteriocins induce membrane leakage and cell death by specifically binding the mannose phophotransferase system (man-PTS) on their target cells. The bacteriocins' cognate immunity proteins that protect the producer cell from its own bacteriocin recognize and bind to the bacteriocin-man-PTS complex and thereby block membrane leakage. In this study, we have determined the three-dimensional structure of the lactococcin A immunity protein (LciA) by the use of nuclear magnetic resonance spectroscopy. LciA forms a four-helix bundle structure with a flexible C-terminal tail. Despite the low degree of sequence similarity between LciA and the pediocin-like immunity proteins, they share the same fold. However, there are certain differences between the structures. The C-terminal helix in LciA is considerably shorter than that observed in the pediocin-like immunity proteins, and the surface potentials of the immunity proteins differ. Truncated variants of LciA in which 6 or 10 of the C-terminal residues were removed yielded a reduced degree of protection, indicating that the unstructured C-terminal tail is important for the functionality of the immunity proteins.
- Department of Biosciences, Section for Biochemistry and Molecular Biology, University of Oslo , P.O. Box 1066, Blindern, 0316 Oslo, Norway.
Organizational Affiliation: