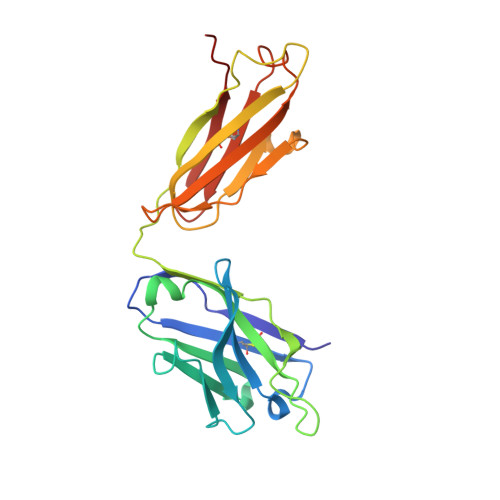
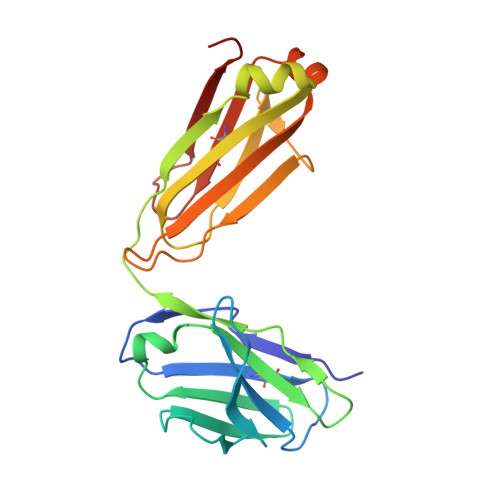
Structure-Based Design of Hepatitis C Virus Vaccines That Elicit Neutralizing Antibody Responses to a Conserved Epitope.
Pierce, B.G., Boucher, E.N., Piepenbrink, K.H., Ejemel, M., Rapp, C.A., Thomas, W.D., Sundberg, E.J., Weng, Z., Wang, Y.(2017) J Virol 91
- PubMed: 28794021 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JVI.01032-17
- Primary Citation Related Structures:
5KZP - PubMed Abstract:
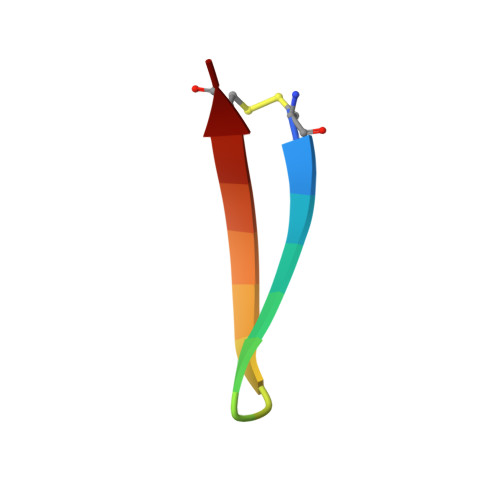
Despite recent advances in therapeutic options, hepatitis C virus (HCV) remains a severe global disease burden, and a vaccine can substantially reduce its incidence. Due to its extremely high sequence variability, HCV can readily escape the immune response; thus, an effective vaccine must target conserved, functionally important epitopes. Using the structure of a broadly neutralizing antibody in complex with a conserved linear epitope from the HCV E2 envelope glycoprotein (residues 412 to 423; epitope I), we performed structure-based design of immunogens to induce antibody responses to this epitope. This resulted in epitope-based immunogens based on a cyclic defensin protein, as well as a bivalent immunogen with two copies of the epitope on the E2 surface. We solved the X-ray structure of a cyclic immunogen in complex with the HCV1 antibody and confirmed preservation of the epitope conformation and the HCV1 interface. Mice vaccinated with our designed immunogens produced robust antibody responses to epitope I, and their serum could neutralize HCV. Notably, the cyclic designs induced greater epitope-specific responses and neutralization than the native peptide epitope. Beyond successfully designing several novel HCV immunogens, this study demonstrates the principle that neutralizing anti-HCV antibodies can be induced by epitope-based, engineered vaccines and provides the basis for further efforts in structure-based design of HCV vaccines. IMPORTANCE Hepatitis C virus is a leading cause of liver disease and liver cancer, with approximately 3% of the world's population infected. To combat this virus, an effective vaccine would have distinct advantages over current therapeutic options, yet experimental vaccines have not been successful to date, due in part to the virus's high sequence variability leading to immune escape. In this study, we rationally designed several vaccine immunogens based on the structure of a conserved epitope that is the target of broadly neutralizing antibodies. In vivo results in mice indicated that these antigens elicited epitope-specific neutralizing antibodies, with various degrees of potency and breadth. These promising results suggest that a rational design approach can be used to generate an effective vaccine for this virus.
- Program in Bioinformatics and Integrative Biology, University of Massachusetts Medical School, Worcester, Massachusetts, USA pierce@umd.edu yang.wang@umassmed.edu.
Organizational Affiliation: