Crystal structure of Mycobacterium tuberculosis hypoxanthine guanine phosphoribosyltransferase in complex with pyrrolidine nucleoside phosphonate
Eng, W.S., Rejman, D., Keough, D.T., Guddat, L.W.To be published.
Experimental Data Snapshot
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Hypoxanthine-guanine phosphoribosyltransferase | 207 | Mycobacterium tuberculosis H37Rv | Mutation(s): 0 Gene Names: hpt, hprT, Rv3624c, MTCY15C10.28 EC: 2.4.2.8 | 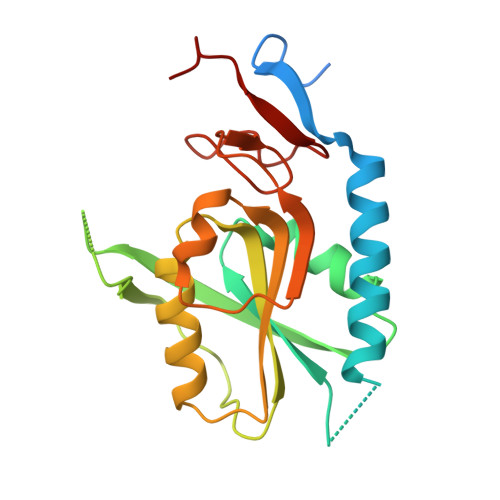 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P9WHQ9 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| YPG Download:Ideal Coordinates CCD File | E [auth A], G [auth B], I [auth C], K [auth D] | [3-[(3~{R},4~{R})-3-(2-azanyl-6-oxidanylidene-1~{H}-purin-9-yl)-4-[(2~{S})-2-oxidanyl-2-phosphono-ethoxy]pyrrolidin-1-y
l]-3-oxidanylidene-propyl]phosphonic acid C14 H22 N6 O10 P2 RBYFDIJTTUISNF-MRTMQBJTSA-N |  | ||
| MG Download:Ideal Coordinates CCD File | F [auth A], H [auth B], J [auth C], L [auth D] | MAGNESIUM ION Mg JLVVSXFLKOJNIY-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 54.523 | α = 90 |
| b = 86.075 | β = 105.95 |
| c = 79.824 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| PDB_EXTRACT | data extraction |
| XDS | data reduction |
| SCALA | data scaling |
| PHASER | phasing |