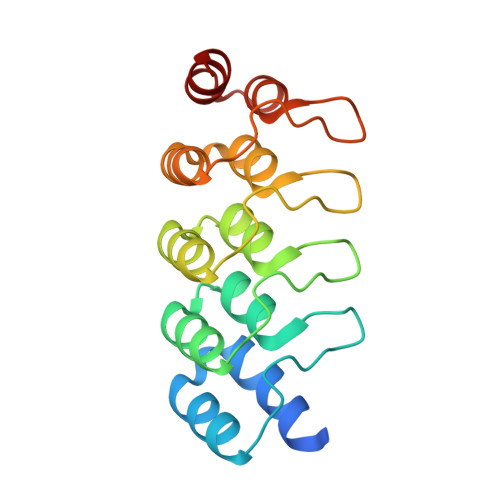
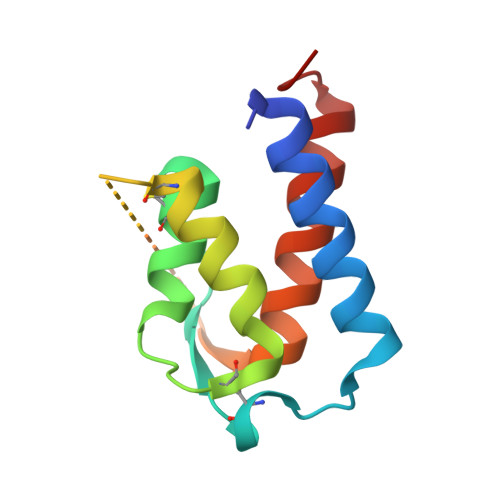
Conformational flexibility of an anti-IL-13 DARPin.
Teplyakov, A., Malia, T.J., Obmolova, G., Jacobs, S.A., O'Neil, K.T., Gilliland, G.L.(2017) Protein Eng Des Sel 30: 31-37
- PubMed: 27881684 Search on PubMed
- DOI: https://doi.org/10.1093/protein/gzw059
- Primary Citation Related Structures:
5KNG, 5KNH - PubMed Abstract:
Designed ankyrin repeat proteins (DARPin ® ) are artificial non-immunoglobulin binding proteins with potential applications as therapeutic molecules. DARPin 6G9 binds interleukin-13 with high affinity and blocks the signaling pathway and as such is promising for the treatment of asthma and other atopic diseases. The crystal structures of DARPin 6G9 in the unbound form and in complex with IL-13 were determined at high resolution. The DARPin competes for the same epitope as the IL-13 receptor chain 13Rα1 but does not interfere with the binding of the other receptor chain, IL-4Rα. Analysis of multiple copies of the DARPin molecule in the crystal indicates the conformational instability in the N-terminal cap that was predicted from molecular dynamics simulations. Comparison of the DARPin structures in the free state and in complex with IL-13 reveals a concerted movement of the ankyrin repeats upon binding resulted in the opening of the binding site. The induced-fit mode of binding employed by DARPin 6G9 is very unusual for DARPins since they were designed as particularly stable and rigid molecules. This finding shows that DARPins can operate by various binding mechanisms and suggests that some flexibility in the scaffold may be an advantage.
- Janssen Research & Development, LLC, 1400 McKean Road, Spring House, PA 19477, USA ateplyak@its.jnj.com.
Organizational Affiliation: