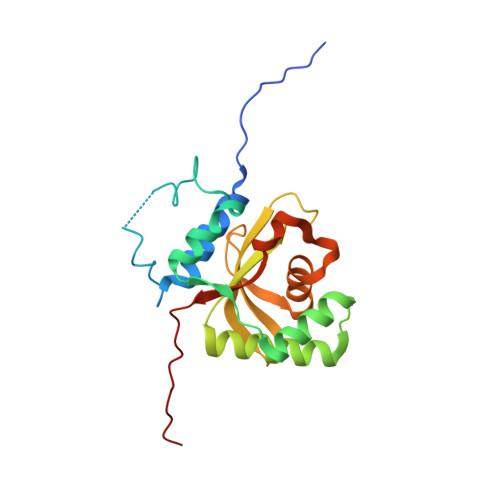
Crystal Structure of the Salmonella Typhimurium Effector GtgE.
Xu, C., Kozlov, G., Wong, K., Gehring, K., Cygler, M.(2016) PLoS One 11: e0166643-e0166643
- PubMed: 27923041 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0166643
- Primary Citation Related Structures:
5KDG - PubMed Abstract:
Salmonella Typhimurium GtgE is an effector protein contributing to the virulence of this pathogen. It was shown to possess highly selective proteolytic activity against a subset of Rab proteins that helps in evasion of Salmonella-containing vacuole (SCV) fusion with lysosomes. Cys45, His151 and Asp169 are essential for proteolytic activity. The structure of a C-terminal fragment GtgE(79-214) indicated the presence of a papain-like fold. Here, we present the structure of GtgE(17-214) containing the fully assembled active site. The design of a proteolytically active and crystallizable GtgE construct was aided by NMR spectroscopy. The protein indeed displays papain-like fold with an assembled Cys-His-Asp catalytic triad. Like the full-length GtgE, the crystallizable construct showed low activity in vitro for its known substrates, Rab32 and Rab29. NMR titration experiments showed at most very weak binding of GtgE to the peptide encompassing the Rab29 cleavage site. In view of the low in vitro activity and poor substrate binding, we postulate that the function of GtgE in vivo as a proteolytic enzyme is dependent on other factor(s), such as a protein partner or interactions with the SCV membrane, which stimulate(s) GtgE activity in vivo.
- Department of Biochemistry, University of Saskatchewan, Saskatoon, Saskatchewan, Canada.
Organizational Affiliation: