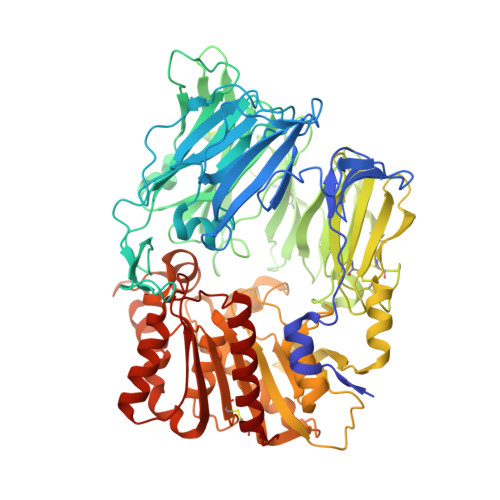
Trelagliptin (SYR-472, Zafatek), Novel Once-Weekly Treatment for Type 2 Diabetes, Inhibits Dipeptidyl Peptidase-4 (DPP-4) via a Non-Covalent Mechanism.
Grimshaw, C.E., Jennings, A., Kamran, R., Ueno, H., Nishigaki, N., Kosaka, T., Tani, A., Sano, H., Kinugawa, Y., Koumura, E., Shi, L., Takeuchi, K.(2016) PLoS One 11: e0157509-e0157509
- PubMed: 27328054 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0157509
- Primary Citation Related Structures:
5KBY - PubMed Abstract:
Trelagliptin (SYR-472), a novel dipeptidyl peptidase-4 inhibitor, shows sustained efficacy by once-weekly dosing in type 2 diabetes patients. In this study, we characterized in vitro properties of trelagliptin, which exhibited approximately 4- and 12-fold more potent inhibition against human dipeptidyl peptidase-4 than alogliptin and sitagliptin, respectively, and >10,000-fold selectivity over related proteases including dipeptidyl peptidase-8 and dipeptidyl peptidase-9. Kinetic analysis revealed reversible, competitive and slow-binding inhibition of dipeptidyl peptidase-4 by trelagliptin (t1/2 for dissociation ≈ 30 minutes). X-ray diffraction data indicated a non-covalent interaction between dipeptidyl peptidase and trelagliptin. Taken together, potent dipeptidyl peptidase inhibition may partially contribute to sustained efficacy of trelagliptin.
- Enzymology and Biophysical Chemistry, Takeda California, Inc., San Diego, California, United States of America.
Organizational Affiliation: