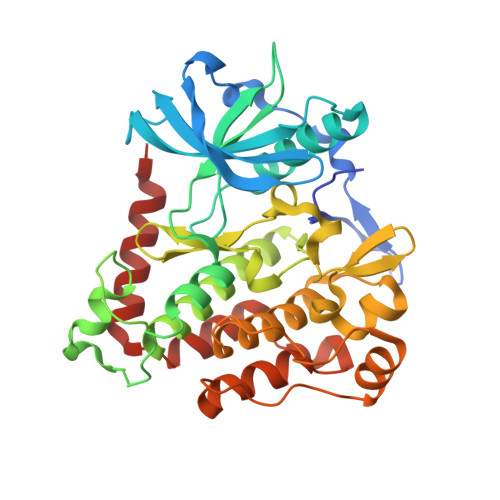
Structural and biochemical studies of the PDGFRA kinase domain
Liang, L., Yan, X.E., Yin, Y., Yun, C.H.(2016) Biochem Biophys Res Commun 477: 667-672
- PubMed: 27349873 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2016.06.117
- Primary Citation Related Structures:
5K5X - PubMed Abstract:
Platelet-derived growth factor receptor α (PDGFRA) is a Type III receptor tyrosine kinase, and this kinase is a target for treatment of gastrointestinal stromal tumors (GIST) as it is frequently mutated in these cancers. Most of the mutations that cause constitutive activation of PDGFRA occur in either the activation loop (A-loop) or in the juxtamembrane (JM) domain, such as the mutations D842V or V561D respectively. Treatment of PDGFRA-mutated GIST with imatinib is successful in some cases, but the D842V mutation is imatinib-resistant. To better understand the mechanism of PDGFRA drug-resistance, we have determined the crystal structure of the PDGFRA kinase domain in the auto-inhibited form, and studied the kinetics of the D842V mutation. Auto-inhibited PDGFRA is stabilized by the JM domain, which inserts into the active site of the kinase. The conserved residue Asp842 makes extensive contacts with several A-loop residues to maintain PDGFRA in the "DFG out" conformation, which stabilizes the kinase in the inactive state and facilitates the binding of imatinib. The D842V mutation would therefore be expected to activate the kinase and hinder the binding of drug through destabilizing the "DFG out" conformation. Furthermore, our kinetic data show that drug resistance in the D842V mutation may also in part result from its increased affinity for ATP. The PDGFRA kinase domain structure we report in this study has potential to facilitate development of new agents which can inhibit this kinase, including both its activating and drug-resistant mutations.
- Institute of Systems Biomedicine, School of Basic Medical Sciences, Peking University Health Science Center, Beijing 100191, PR China; Department of Pathology, School of Basic Medical Sciences, Peking University Health Science Center, Beijing 100191, PR China.
Organizational Affiliation: