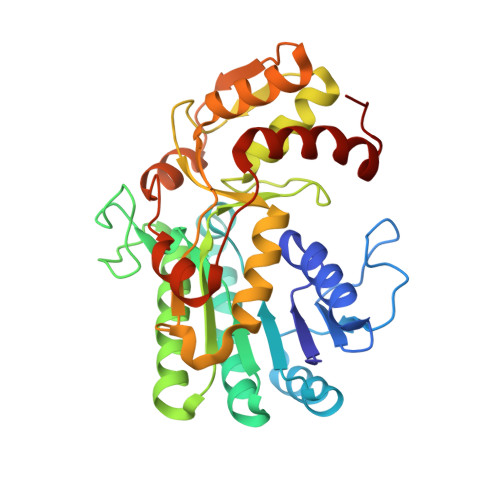
Structure and function of L-threonine-3-dehydrogenase from the parasitic protozoan Trypanosoma brucei revealed by X-ray crystallography and geometric simulations.
Adjogatse, E., Erskine, P., Wells, S.A., Kelly, J.M., Wilden, J.D., Chan, A.W.E., Selwood, D., Coker, A., Wood, S., Cooper, J.B.(2018) Acta Crystallogr D Struct Biol 74: 861-876
- PubMed: 30198897 Search on PubMed
- DOI: https://doi.org/10.1107/S2059798318009208
- Primary Citation Related Structures:
5K4Q, 5K4T, 5K4U, 5K4V, 5K4W, 5K4Y, 5K50, 5L9A, 5LC1 - PubMed Abstract:
Two of the world's most neglected tropical diseases, human African trypanosomiasis (HAT) and Chagas disease, are caused by protozoan parasites of the genus Trypanosoma. These organisms possess specialized metabolic pathways, frequently distinct from those in humans, which have potential to be exploited as novel drug targets. This study elucidates the structure and function of L-threonine-3-dehydrogenase (TDH) from T. brucei, the causative pathogen of HAT. TDH is a key enzyme in the metabolism of L-threonine, and an inhibitor of TDH has been shown to have trypanocidal activity in the procyclic form of T. brucei. TDH is a nonfunctional pseudogene in humans, suggesting that it may be possible to rationally design safe and specific therapies for trypanosomiasis by targeting this parasite enzyme. As an initial step, the TDH gene from T. brucei was expressed and the three-dimensional structure of the enzyme was solved by X-ray crystallography. In multiple crystallographic structures, T. brucei TDH is revealed to be a dimeric short-chain dehydrogenase that displays a considerable degree of conformational variation in its ligand-binding regions. Geometric simulations of the structure have provided insight into the dynamic behaviour of this enzyme. Furthermore, structures of TDH bound to its natural substrates and known inhibitors have been determined, giving an indication of the mechanism of catalysis of the enzyme. Collectively, these results provide vital details for future drug design to target TDH or related enzymes.
- Laboratory for Protein Crystallography, Division of Medicine, University College London, Gower Street, London WC1E 7HT, England.
Organizational Affiliation: